Conserved active site cysteine residue of archaeal THI4 homolog is essential for thiamine biosynthesis in Haloferax volcanii
- PMID: 25348237
- PMCID: PMC4215014
- DOI: 10.1186/s12866-014-0260-0
Conserved active site cysteine residue of archaeal THI4 homolog is essential for thiamine biosynthesis in Haloferax volcanii
Abstract
Background: Thiamine (vitamin B1) is synthesized de novo by certain yeast, fungi, plants, protozoans, bacteria and archaea. The pathway of thiamine biosynthesis by archaea is poorly understood, particularly the route of sulfur relay to form the thiazole ring. Archaea harbor structural homologs of both the bacterial (ThiS-ThiF) and eukaryotic (THI4) proteins that mobilize sulfur to thiazole ring precursors by distinct mechanisms.
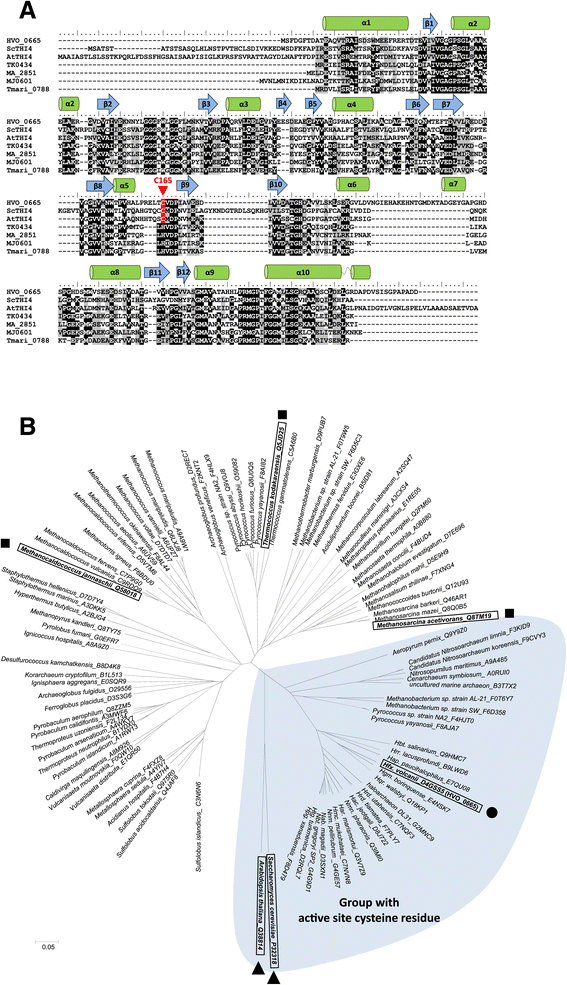
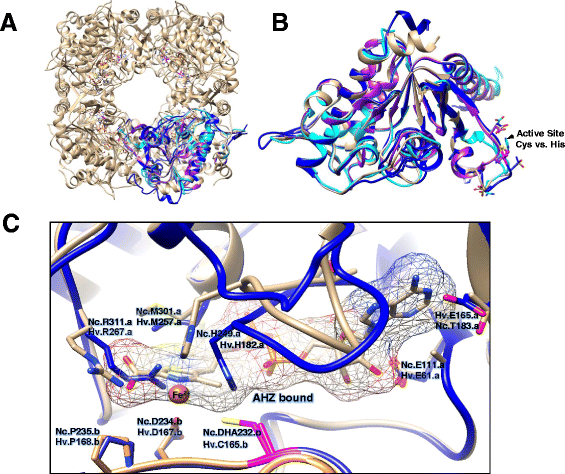
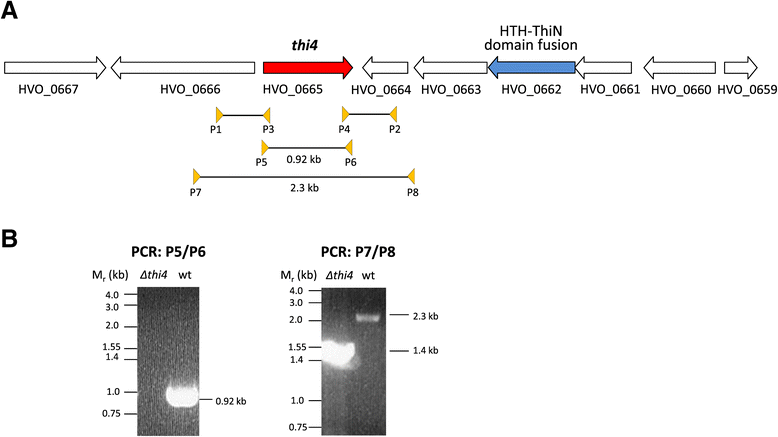
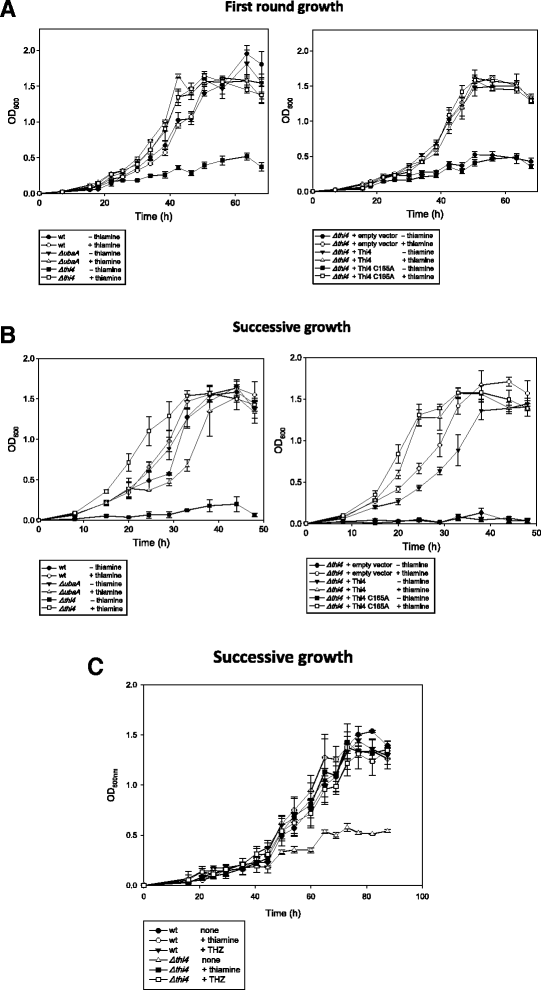
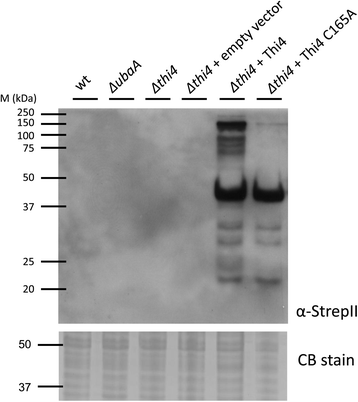
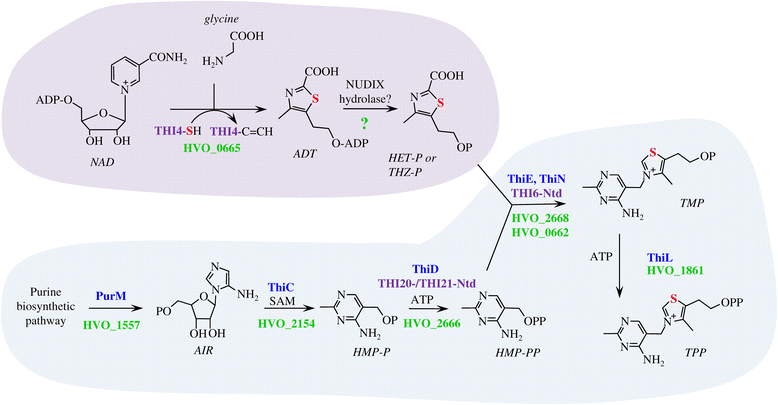
Results: Based on comparative genome analysis, halophilic archaea are predicted to synthesize the pyrimidine moiety of thiamine by the bacterial pathway, initially suggesting that also a bacterial ThiS-ThiF type mechanism for synthesis of the thiazole ring is used in which the sulfur carrier ThiS is first activated by ThiF-catalyzed adenylation. The only ThiF homolog of Haloferax volcanii (UbaA) was deleted but this had no effect on growth in the absence of thiamine. Usage of the eukaryotic THI4-type sulfur relay was initially considered less likely for thiamine biosynthesis in archaea, since the active-site cysteine residue of yeast THI4p that donates the sulfur to the thiazole ring by a suicide mechanism is replaced by a histidine residue in many archaeal THI4 homologs and these are described as D-ribose-1,5-bisphosphate isomerases. The THI4 homolog of the halophilic archaea, including Hfx. volcanii (HVO_0665, HvThi4) was found to differ from that of methanogens and thermococci by having a cysteine residue (Cys165) corresponding to the conserved active site cysteine of yeast THI4p (Cys205). Deletion of HVO_0665 generated a thiamine auxotroph that was trans-complemented by a wild-type copy of HVO_0665, but not the modified gene encoding an HvThi4 C165A variant.
Conclusions: Based on our results, we conclude that the archaeon Hfx. volcanii uses a yeast THI4-type mechanism for sulfur relay to form the thiazole ring of thiamine. We extend this finding to a relatively large group of archaea, including haloarchaea, ammonium oxidizing archaea, and some methanogen and Pyrococcus species, by observing that these organisms code for THI4 homologs that have a conserved active site cysteine residue which is likely used in thiamine biosynthesis. Thus, archaeal members of IPR002922 THI4 family that have a conserved cysteine active site should be reexamined for a function in thiamine biosynthesis.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

