Cancer in silico drug discovery: a systems biology tool for identifying candidate drugs to target specific molecular tumor subtypes
- PMID: 25349306
- PMCID: PMC4341901
- DOI: 10.1158/1535-7163.MCT-14-0260
Cancer in silico drug discovery: a systems biology tool for identifying candidate drugs to target specific molecular tumor subtypes
Abstract
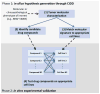
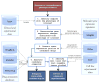
Large-scale cancer datasets such as The Cancer Genome Atlas (TCGA) allow researchers to profile tumors based on a wide range of clinical and molecular characteristics. Subsequently, TCGA-derived gene expression profiles can be analyzed with the Connectivity Map (CMap) to find candidate drugs to target tumors with specific clinical phenotypes or molecular characteristics. This represents a powerful computational approach for candidate drug identification, but due to the complexity of TCGA and technology differences between CMap and TCGA experiments, such analyses are challenging to conduct and reproduce. We present Cancer in silico Drug Discovery (CiDD; scheet.org/software), a computational drug discovery platform that addresses these challenges. CiDD integrates data from TCGA, CMap, and Cancer Cell Line Encyclopedia (CCLE) to perform computational drug discovery experiments, generating hypotheses for the following three general problems: (i) determining whether specific clinical phenotypes or molecular characteristics are associated with unique gene expression signatures; (ii) finding candidate drugs to repress these expression signatures; and (iii) identifying cell lines that resemble the tumors being studied for subsequent in vitro experiments. The primary input to CiDD is a clinical or molecular characteristic. The output is a biologically annotated list of candidate drugs and a list of cell lines for in vitro experimentation. We applied CiDD to identify candidate drugs to treat colorectal cancers harboring mutations in BRAF. CiDD identified EGFR and proteasome inhibitors, while proposing five cell lines for in vitro testing. CiDD facilitates phenotype-driven, systematic drug discovery based on clinical and molecular data from TCGA.
©2014 American Association for Cancer Research.
Conflict of interest statement
Figures

Similar articles
-
Can a Liquid Biopsy Detect Circulating Tumor DNA With Low-passage Whole-genome Sequencing in Patients With a Sarcoma? A Pilot Evaluation.Clin Orthop Relat Res. 2025 Jan 1;483(1):39-48. doi: 10.1097/CORR.0000000000003161. Epub 2024 Jun 21. Clin Orthop Relat Res. 2025. PMID: 38905450
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
EORTC guidelines for the use of erythropoietic proteins in anaemic patients with cancer: 2006 update.Eur J Cancer. 2007 Jan;43(2):258-70. doi: 10.1016/j.ejca.2006.10.014. Epub 2006 Dec 19. Eur J Cancer. 2007. PMID: 17182241
Cited by
-
Nanoinformatics and Personalized Medicine: An Advanced Cumulative Approach for Cancer Management.Curr Med Chem. 2023;30(3):271-285. doi: 10.2174/0929867329666220610090405. Curr Med Chem. 2023. PMID: 35692148 Review.
-
Proteomic Profiling of BRAFV600E Mutant Colon Cancer Cells Reveals the Involvement of Nucleophosmin/c-Myc Axis in Modulating the Response and Resistance to BRAF Inhibition by Vemurafenib.Int J Mol Sci. 2021 Jun 8;22(12):6174. doi: 10.3390/ijms22126174. Int J Mol Sci. 2021. PMID: 34201061 Free PMC article.
-
paraGSEA: a scalable approach for large-scale gene expression profiling.Nucleic Acids Res. 2017 Sep 29;45(17):e155. doi: 10.1093/nar/gkx679. Nucleic Acids Res. 2017. PMID: 28973463 Free PMC article.
-
Predicting Novel Therapies and Targets: Regulation of Notch3 by the Bromodomain Protein BRD4.Mol Cancer Ther. 2019 Feb;18(2):421-436. doi: 10.1158/1535-7163.MCT-18-0365. Epub 2018 Nov 12. Mol Cancer Ther. 2019. PMID: 30420565 Free PMC article.
-
Mathematical Modeling for the Assessment of Public Policies in the Cancer Health-Care System Implemented for the Colombian Case.Int J Environ Res Public Health. 2023 Sep 11;20(18):6740. doi: 10.3390/ijerph20186740. Int J Environ Res Public Health. 2023. PMID: 37754600 Free PMC article.
References
-
- Lievre A, Bachet J-B, Boige V, Cayre A, Le Corre D, Buc E, et al. KRAS Mutations As an Independent Prognostic Factor in Patients With Advanced Colorectal Cancer Treated With Cetuximab. J Clin Oncol. 2008 Jan 20;26:374–9. - PubMed
-
- Prahallad A, Sun C, Huang S, Di Nicolantonio F, Salazar R, Zecchin D, et al. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature. 2012 Jan 26;483:100–3. - PubMed
-
- McDermott U, Settleman J. Personalized Cancer Therapy With Selective Kinase Inhibitors: An Emerging Paradigm in Medical Oncology. J Clin Oncol. 2009 Oct 26;27:5650–9. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

