Poly-dA:dT tracts form an in vivo nucleosomal turnstile
- PMID: 25353956
- PMCID: PMC4212969
- DOI: 10.1371/journal.pone.0110479
Poly-dA:dT tracts form an in vivo nucleosomal turnstile
Abstract
Nucleosomes regulate many DNA-dependent processes by controlling the accessibility of DNA, and DNA sequences such as the poly-dA:dT element are known to affect nucleosome binding. We demonstrate that poly-dA:dT tracts form an asymmetric barrier to nucleosome movement in vivo, mediated by ATP-dependent chromatin remodelers. We theorize that nucleosome transit over poly-A elements is more energetically favourable in one direction, leading to an asymmetric arrangement of nucleosomes around these sequences. We demonstrate that different arrangements of poly-A and poly-T tracts result in very different outcomes for nucleosome occupancy in yeast, mouse, and human, and show that yeast takes advantage of this phenomenon in its promoter architecture.
Conflict of interest statement
Figures

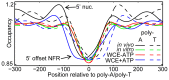

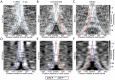
References
-
- Hampson S, Kibler D, Baldi P (2002) Distribution patterns of over-represented k-mers in non-coding yeast DNA. Bioinformatics 18: 513–528. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

