Structural analysis of a novel small molecule ligand bound to the CXCL12 chemokine
- PMID: 25356720
- PMCID: PMC4255719
- DOI: 10.1021/jm501194p
Structural analysis of a novel small molecule ligand bound to the CXCL12 chemokine
Abstract
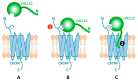
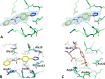
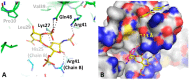
CXCL12 binds to CXCR4, promoting both chemotaxis of lymphocytes and metastasis of cancer cells. We previously identified small molecule ligands that bind CXCL12 and block CXCR4-mediated chemotaxis. We now report a 1.9 Å resolution X-ray structure of CXCL12 bound by such a molecule at a site normally bound by sY21 of CXCR4. The complex structure reveals binding hot spots for future inhibitor design and suggests a new approach to targeting CXCL12-CXCR4 signaling in drug discovery.
Figures
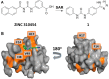

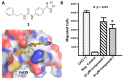
Similar articles
-
Fragment-based optimization of small molecule CXCL12 inhibitors for antagonizing the CXCL12/CXCR4 interaction.Curr Top Med Chem. 2012;12(24):2727-40. doi: 10.2174/1568026611212240003. Curr Top Med Chem. 2012. PMID: 23368099 Free PMC article.
-
Targeting SDF-1/CXCL12 with a ligand that prevents activation of CXCR4 through structure-based drug design.J Am Chem Soc. 2010 Jun 2;132(21):7242-3. doi: 10.1021/ja1002263. J Am Chem Soc. 2010. PMID: 20459090 Free PMC article.
-
Interaction of chemokine receptor CXCR4 in monomeric and dimeric state with its endogenous ligand CXCL12: coarse-grained simulations identify differences.J Biomol Struct Dyn. 2017 Feb;35(2):399-412. doi: 10.1080/07391102.2016.1145142. Epub 2016 Jul 11. J Biomol Struct Dyn. 2017. PMID: 26813575
-
The good and bad faces of the CXCR4 chemokine receptor.Int J Biochem Cell Biol. 2018 Feb;95:121-131. doi: 10.1016/j.biocel.2017.12.018. Epub 2017 Dec 27. Int J Biochem Cell Biol. 2018. PMID: 29288743 Review.
-
Biological/pathological functions of the CXCL12/CXCR4/CXCR7 axes in the pathogenesis of bladder cancer.Int J Clin Oncol. 2017 Dec;22(6):991-1000. doi: 10.1007/s10147-017-1187-x. Epub 2017 Oct 11. Int J Clin Oncol. 2017. PMID: 29022185 Review.
Cited by
-
Crystallographic Structure of Truncated CCL21 and the Putative Sulfotyrosine-Binding Site.Biochemistry. 2016 Oct 11;55(40):5746-5753. doi: 10.1021/acs.biochem.6b00304. Epub 2016 Sep 26. Biochemistry. 2016. PMID: 27617343 Free PMC article.
-
Modeling the SDF-1/CXCR4 protein using advanced artificial intelligence and antagonist screening for Japanese anchovy.Front Physiol. 2024 Feb 2;15:1349119. doi: 10.3389/fphys.2024.1349119. eCollection 2024. Front Physiol. 2024. PMID: 38370015 Free PMC article.
-
C-X-C motif chemokine ligand 12-C-X-C chemokine receptor type 4 signaling axis in cancer and the development of chemotherapeutic molecules.Tzu Chi Med J. 2024 May 27;36(3):231-239. doi: 10.4103/tcmj.tcmj_52_24. eCollection 2024 Jul-Sep. Tzu Chi Med J. 2024. PMID: 38993827 Free PMC article. Review.
-
Structural Insights into Molecular Recognition by Human Chemokine CCL19.Biochemistry. 2022 Mar 1;61(5):311-318. doi: 10.1021/acs.biochem.1c00759. Epub 2022 Feb 14. Biochemistry. 2022. PMID: 35156805 Free PMC article.
-
Structural Analysis of Chemokine Receptor-Ligand Interactions.J Med Chem. 2017 Jun 22;60(12):4735-4779. doi: 10.1021/acs.jmedchem.6b01309. Epub 2017 Mar 10. J Med Chem. 2017. PMID: 28165741 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

