Rad51 and Rad52 are involved in homologous recombination of replicating herpes simplex virus DNA
- PMID: 25365323
- PMCID: PMC4218770
- DOI: 10.1371/journal.pone.0111584
Rad51 and Rad52 are involved in homologous recombination of replicating herpes simplex virus DNA
Abstract
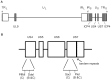
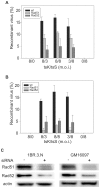
Replication of herpes simplex virus 1 is coupled to recombination, but the molecular mechanisms underlying this process are poorly characterized. The role of Rad51 and Rad52 recombinases in viral recombination was examined in human fibroblast cells 1BR.3.N (wild type) and in GM16097 with replication defects caused by mutations in DNA ligase I. Intermolecular recombination between viruses, tsS and tsK, harboring genetic markers gave rise to ∼17% recombinants in both cell lines. Knock-down of Rad51 and Rad52 by siRNA reduced production of recombinants to 11% and 5%, respectively, in wild type cells and to 3% and 5%, respectively, in GM16097 cells. The results indicate a specific role for Rad51 and Rad52 in recombination of replicating herpes simplex virus 1 DNA. Mixed infections using clinical isolates with restriction enzyme polymorphisms in the US4 and US7 genes revealed recombination frequencies of 0.7%/kbp in wild type cells and 4%/kbp in GM16097 cells. Finally, tandem repeats in the US7 gene remained stable upon serial passage, indicating a high fidelity of recombination in infected cells.
Conflict of interest statement
Figures
Similar articles
-
Rad51-Rad52 mediated maintenance of centromeric chromatin in Candida albicans.PLoS Genet. 2014 Apr 24;10(4):e1004344. doi: 10.1371/journal.pgen.1004344. eCollection 2014 Apr. PLoS Genet. 2014. PMID: 24762765 Free PMC article.
-
Recovery of deficient homologous recombination in Brca2-depleted mouse cells by wild-type Rad51 expression.DNA Repair (Amst). 2009 Feb 1;8(2):170-81. doi: 10.1016/j.dnarep.2008.10.002. Epub 2008 Nov 18. DNA Repair (Amst). 2009. PMID: 18992372
-
Dynamic regulatory interactions of rad51, rad52, and replication protein-a in recombination intermediates.J Mol Biol. 2009 Jul 3;390(1):45-55. doi: 10.1016/j.jmb.2009.05.009. Epub 2009 May 13. J Mol Biol. 2009. PMID: 19445949
-
Mechanism of homologous recombination and implications for aging-related deletions in mitochondrial DNA.Microbiol Mol Biol Rev. 2013 Sep;77(3):476-96. doi: 10.1128/MMBR.00007-13. Microbiol Mol Biol Rev. 2013. PMID: 24006472 Free PMC article. Review.
-
Non-Recombinogenic Functions of Rad51, BRCA2, and Rad52 in DNA Damage Tolerance.Genes (Basel). 2021 Sep 29;12(10):1550. doi: 10.3390/genes12101550. Genes (Basel). 2021. PMID: 34680945 Free PMC article. Review.
Cited by
-
Transcriptional analysis reveals the suppression of RAD51 and disruption of the homologous recombination pathway during PEDV infection in IPEC-J2 cells.Virol J. 2024 Dec 28;21(1):337. doi: 10.1186/s12985-024-02611-8. Virol J. 2024. PMID: 39731192 Free PMC article.
-
Rapid poxvirus engineering using CRISPR/Cas9 as a selection tool.Commun Biol. 2020 Nov 3;3(1):643. doi: 10.1038/s42003-020-01374-6. Commun Biol. 2020. PMID: 33144673 Free PMC article.
-
Herpes Simplex Virus 1 Manipulates Host Cell Antiviral and Proviral DNA Damage Responses.mBio. 2021 Feb 9;12(1):e03552-20. doi: 10.1128/mBio.03552-20. mBio. 2021. PMID: 33563816 Free PMC article.
-
Localization of Double-Strand Break Repair Proteins to Viral Replication Compartments following Lytic Reactivation of Kaposi's Sarcoma-Associated Herpesvirus.J Virol. 2017 Oct 27;91(22):e00930-17. doi: 10.1128/JVI.00930-17. Print 2017 Nov 15. J Virol. 2017. PMID: 28855246 Free PMC article.
-
Whole genome sequencing of Herpes Simplex Virus 1 directly from human cerebrospinal fluid reveals selective constraints in neurotropic viruses.Virus Evol. 2020 Feb 20;6(1):veaa012. doi: 10.1093/ve/veaa012. eCollection 2020 Jan. Virus Evol. 2020. PMID: 32099667 Free PMC article.
References
-
- Bowden R, Sakaoka H, Donnelly P, Ward R (2004) High recombination rate in herpes simplex virus type 1 natural populations suggests significant co-infection. Infection, genetics and evolution: journal of molecular epidemiology and evolutionary genetics in infectious diseases 4: 115–123. - PubMed
-
- Wilkinson DE, Weller SK (2003) The role of DNA recombination in herpes simplex virus DNA replication. IUBMB Life 55: 451–458. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

