Coordination of insulin and Notch pathway activities by microRNA miR-305 mediates adaptive homeostasis in the intestinal stem cells of the Drosophila gut
- PMID: 25367037
- PMCID: PMC4215186
- DOI: 10.1101/gad.241588.114
Coordination of insulin and Notch pathway activities by microRNA miR-305 mediates adaptive homeostasis in the intestinal stem cells of the Drosophila gut
Abstract
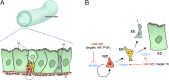
Homeostasis of the intestine is maintained by dynamic regulation of a pool of intestinal stem cells. The balance between stem cell self-renewal and differentiation is regulated by the Notch and insulin signaling pathways. Dependence on the insulin pathway places the stem cell pool under nutritional control, allowing gut homeostasis to adapt to environmental conditions. Here we present evidence that miR-305 is required for adaptive homeostasis of the gut. miR-305 regulates the Notch and insulin pathways in the intestinal stem cells. Notably, miR-305 expression in the stem cells is itself under nutritional control via the insulin pathway. This link places regulation of Notch pathway activity under nutritional control. These findings provide a mechanism through which the insulin pathway controls the balance between stem cell self-renewal and differentiation that is required for adaptive homeostasis in the gut in response to changing environmental conditions.
Keywords: Notch; insulin; microRNA; stem cell.
© 2014 Foronda et al.; Published by Cold Spring Harbor Laboratory Press.
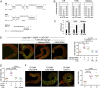
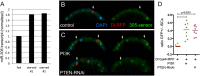
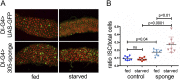
Figures
Similar articles
-
InR and Pi3K maintain intestinal homeostasis through STAT/EGFR and Notch signaling in enteroblasts.J Cell Biochem. 2024 Jun;125(6):e30545. doi: 10.1002/jcb.30545. Epub 2024 Mar 4. J Cell Biochem. 2024. PMID: 38436545
-
Paracrine Wingless signalling controls self-renewal of Drosophila intestinal stem cells.Nature. 2008 Oct 23;455(7216):1119-23. doi: 10.1038/nature07329. Epub 2008 Sep 21. Nature. 2008. PMID: 18806781
-
Opposing activities of Notch and Wnt signaling regulate intestinal stem cells and gut homeostasis.Cell Rep. 2015 Apr 7;11(1):33-42. doi: 10.1016/j.celrep.2015.03.007. Epub 2015 Mar 26. Cell Rep. 2015. PMID: 25818302 Free PMC article.
-
Intestinal stem cells in the adult Drosophila midgut.Exp Cell Res. 2011 Nov 15;317(19):2780-8. doi: 10.1016/j.yexcr.2011.07.020. Epub 2011 Aug 11. Exp Cell Res. 2011. PMID: 21856297 Free PMC article. Review.
-
Crosstalk between Wnt and Notch signaling in intestinal epithelial cell fate decision.J Gastroenterol. 2007 Sep;42(9):705-10. doi: 10.1007/s00535-007-2087-z. Epub 2007 Sep 25. J Gastroenterol. 2007. PMID: 17876539 Review.
Cited by
-
Identification and Characterization of Small RNA Markers of Age in the Blow Fly Cochliomyia macellaria (Fabricius) (Diptera: Calliphoridae).Insects. 2022 Oct 18;13(10):948. doi: 10.3390/insects13100948. Insects. 2022. PMID: 36292896 Free PMC article.
-
The role of micro RNAs (miRNAs) in the regulation of Drosophila melanogaster's innate immunity.Fly (Austin). 2022 Dec;16(1):382-396. doi: 10.1080/19336934.2022.2149204. Fly (Austin). 2022. PMID: 36412256 Free PMC article. Review.
-
A stress-responsive miRNA regulates BMP signaling to maintain tissue homeostasis.Proc Natl Acad Sci U S A. 2021 May 25;118(21):e2022583118. doi: 10.1073/pnas.2022583118. Proc Natl Acad Sci U S A. 2021. PMID: 34016750 Free PMC article.
-
The transcription factor Hey and nuclear lamins specify and maintain cell identity.Elife. 2019 Jul 16;8:e44745. doi: 10.7554/eLife.44745. Elife. 2019. PMID: 31310235 Free PMC article.
-
Control of Drosophila Type I and Type II central brain neuroblast proliferation by bantam microRNA.Development. 2015 Nov 1;142(21):3713-20. doi: 10.1242/dev.127209. Epub 2015 Sep 22. Development. 2015. PMID: 26395494 Free PMC article.
References
-
- Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM. 2003. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the pro-apoptotic gene hid in Drosophila. Cell 113: 25–36 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
