Mitochondrial targets for pharmacological intervention in human disease
- PMID: 25367773
- PMCID: PMC4286170
- DOI: 10.1021/pr500813f
Mitochondrial targets for pharmacological intervention in human disease
Abstract
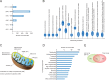
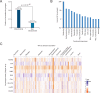
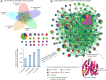
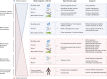
Over the past several years, mitochondrial dysfunction has been linked to an increasing number of human illnesses, making mitochondrial proteins (MPs) an ever more appealing target for therapeutic intervention. With 20% of the mitochondrial proteome (312 of an estimated 1500 MPs) having known interactions with small molecules, MPs appear to be highly targetable. Yet, despite these targeted proteins functioning in a range of biological processes (including induction of apoptosis, calcium homeostasis, and metabolism), very few of the compounds targeting MPs find clinical use. Recent work has greatly expanded the number of proteins known to localize to the mitochondria and has generated a considerable increase in MP 3D structures available in public databases, allowing experimental screening and in silico prediction of mitochondrial drug targets on an unprecedented scale. Here, we summarize the current literature on clinically active drugs that target MPs, with a focus on how existing drug targets are distributed across biochemical pathways and organelle substructures. Also, we examine current strategies for mitochondrial drug discovery, focusing on genetic, proteomic, and chemogenomic assays, and relevant model systems. As cell models and screening techniques improve, MPs appear poised to emerge as relevant targets for a wide range of complex human diseases, an eventuality that can be expedited through systematic analysis of MP function.
Keywords: Drug−protein interactions; human disease; mitochondria; model system; network; pathways; pharmacological target; protein complex; small molecules; systems biology.
Figures



References
-
- Vlasblom J.; Jin K.; Kassir S.; Babu M. Exploring mitochondrial system properties of neurodegenerative diseases through interactome mapping. J. Proteomics 2014, 100, 8–24. - PubMed
-
- Hoppins S. The regulation of mitochondrial dynamics. Curr. Opin. Cell Biol. 2014, 29C, 46–52. - PubMed
-
- Kornmann B. The molecular hug between the ER and the mitochondria. Curr. Opin. Cell Biol. 2013, 25, 443–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

