The Porphyromonas gingivalis ferric uptake regulator orthologue binds hemin and regulates hemin-responsive biofilm development
- PMID: 25375181
- PMCID: PMC4222909
- DOI: 10.1371/journal.pone.0111168
The Porphyromonas gingivalis ferric uptake regulator orthologue binds hemin and regulates hemin-responsive biofilm development
Abstract
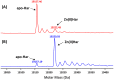
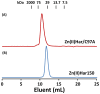
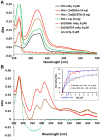
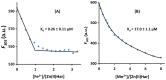
Porphyromonas gingivalis is a Gram-negative pathogen associated with the biofilm-mediated disease chronic periodontitis. P. gingivalis biofilm formation is dependent on environmental heme for which P. gingivalis has an obligate requirement as it is unable to synthesize protoporphyrin IX de novo, hence P. gingivalis transports iron and heme liberated from the human host. Homeostasis of a variety of transition metal ions is often mediated in Gram-negative bacteria at the transcriptional level by members of the Ferric Uptake Regulator (Fur) superfamily. P. gingivalis has a single predicted Fur superfamily orthologue which we have designated Har (heme associated regulator). Recombinant Har formed dimers in the presence of Zn2+ and bound one hemin molecule per monomer with high affinity (Kd of 0.23 µM). The binding of hemin resulted in conformational changes of Zn(II)Har and residue 97Cys was involved in hemin binding as part of a predicted -97C-98P-99L- hemin binding motif. The expression of 35 genes was down-regulated and 9 up-regulated in a Har mutant (ECR455) relative to wild-type. Twenty six of the down-regulated genes were previously found to be up-regulated in P. gingivalis grown as a biofilm and 11 were up-regulated under hemin limitation. A truncated Zn(II)Har bound the promoter region of dnaA (PGN_0001), one of the up-regulated genes in the ECR455 mutant. This binding decreased as hemin concentration increased which was consistent with gene expression being regulated by hemin availability. ECR455 formed significantly less biofilm than the wild-type and unlike wild-type biofilm formation was independent of hemin availability. P. gingivalis possesses a hemin-binding Fur orthologue that regulates hemin-dependent biofilm formation.
Conflict of interest statement
Figures




Similar articles
-
Porphyromonas gingivalis PgFur Is a Member of a Novel Fur Subfamily With Non-canonical Function.Front Cell Infect Microbiol. 2019 Jul 1;9:233. doi: 10.3389/fcimb.2019.00233. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 31312617 Free PMC article.
-
SigCH, an extracytoplasmic function sigma factor of Porphyromonas gingivalis regulates the expression of cdhR and hmuYR.Anaerobe. 2017 Feb;43:82-90. doi: 10.1016/j.anaerobe.2016.12.006. Epub 2016 Dec 8. Anaerobe. 2017. PMID: 27940243
-
Regulation of hemin and iron transport in Porphyromonas gingivalis.Adv Dent Res. 1995 Feb;9(1):41-7. doi: 10.1177/08959374950090010801. Adv Dent Res. 1995. PMID: 7669213 Review.
-
Characterization and expression of HmuR, a TonB-dependent hemoglobin receptor of Porphyromonas gingivalis.J Bacteriol. 2000 Oct;182(20):5737-48. doi: 10.1128/JB.182.20.5737-5748.2000. J Bacteriol. 2000. PMID: 11004172 Free PMC article.
-
Iron and heme utilization in Porphyromonas gingivalis.FEMS Microbiol Rev. 2005 Jan;29(1):119-44. doi: 10.1016/j.femsre.2004.09.001. FEMS Microbiol Rev. 2005. PMID: 15652979 Review.
Cited by
-
New approaches to combat Porphyromonas gingivalis biofilms.J Oral Microbiol. 2017 Mar 15;9(1):1300366. doi: 10.1080/20002297.2017.1300366. eCollection 2017. J Oral Microbiol. 2017. PMID: 28473880 Free PMC article. Review.
-
Exploring heme and iron acquisition strategies of Porphyromonas gingivalis-current facts and hypotheses.FEMS Microbiol Rev. 2025 Jan 14;49:fuaf019. doi: 10.1093/femsre/fuaf019. FEMS Microbiol Rev. 2025. PMID: 40343779 Free PMC article. Review.
-
Comparative Genomics of the Genus Porphyromonas Identifies Adaptations for Heme Synthesis within the Prevalent Canine Oral Species Porphyromonas cangingivalis.Genome Biol Evol. 2015 Nov 13;7(12):3397-413. doi: 10.1093/gbe/evv220. Genome Biol Evol. 2015. PMID: 26568374 Free PMC article.
-
PgFur participates differentially in expression of virulence factors in more virulent A7436 and less virulent ATCC 33277 Porphyromonas gingivalis strains.BMC Microbiol. 2019 Jun 11;19(1):127. doi: 10.1186/s12866-019-1511-x. BMC Microbiol. 2019. PMID: 31185896 Free PMC article.
-
The Porphyromonas gingivalis ferric uptake regulator orthologue does not regulate iron homeostasis.Genom Data. 2015 Jun 11;5:167-8. doi: 10.1016/j.gdata.2015.05.042. eCollection 2015 Sep. Genom Data. 2015. PMID: 26484248 Free PMC article.
References
-
- Loesche WJ, Syed SA, Morrison EC, Laughon B, Grossman NS (1981) Treatment of periodontal infections due to anaerobic bacteria with short-term treatment with metronidazole. J Clin Periodontol 8: 29–44. - PubMed
-
- Slots J (1977) Microflora in the healthy gingival sulcus in man. Scand J Dent Res 85: 247–254. - PubMed
-
- Spiegel CA, Hayduk SE, Minah GE, Krywolap GN (1979) Black-pigmented Bacteroides from clinically characterized periodontal sites. J Periodontal Res 14: 376–382. - PubMed
-
- Van Dyke TE, Offenbacher S, Place D, Dowell VR, Jones J (1988) Refractory periodontitis: mixed infection with Bacteroides gingivalis and other unusual Bacteroides species. A case report. J Periodontol 59: 184–189. - PubMed
-
- White D, Mayrand D (1981) Association of oral Bacteroides with gingivitis and adult periodontitis. J Periodontal Res 16: 259–265. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

