diArk--the database for eukaryotic genome and transcriptome assemblies in 2014
- PMID: 25378341
- PMCID: PMC4384042
- DOI: 10.1093/nar/gku990
diArk--the database for eukaryotic genome and transcriptome assemblies in 2014
Abstract
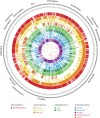
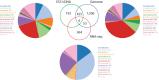
Eukaryotic genomes are the basis for understanding the complexity of life from populations to the molecular level. Recent technological innovations have revolutionized the speed of data generation enabling the sequencing of eukaryotic genomes and transcriptomes within days. The database diArk (http://www.diark.org) has been developed with the aim to provide access to all available assembled genomes and transcriptomes. In September 2014, diArk contains about 2600 eukaryotes with 6000 genome and transcriptome assemblies, of which 22% are not available via NCBI/ENA/DDBJ. Several indicators for the quality of the assemblies are provided to facilitate their comparison for selecting the most appropriate dataset for further studies. diArk has a user-friendly web interface with extensive options for filtering and browsing the sequenced eukaryotes. In this new version of the database we have also integrated species, for which transcriptome assemblies are available, and we provide more analyses of assemblies.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

References
-
- Daetwyler H.D., Capitan A., Pausch H., Stothard P., van Binsbergen R., Brøndum R.F., Liao X., Djari A., Rodriguez S.C., Grohs C., et al. Whole-genome sequencing of 234 bulls facilitates mapping of monogenic and complex traits in cattle. Nat. Genet. 2014;46:858–865. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

