Nutrient-sensing nuclear receptors coordinate autophagy
- PMID: 25383539
- PMCID: PMC4267857
- DOI: 10.1038/nature13961
Nutrient-sensing nuclear receptors coordinate autophagy
Abstract
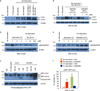
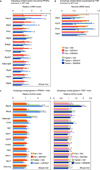
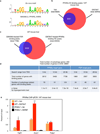
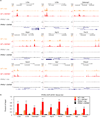
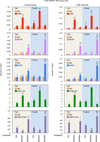
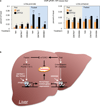
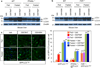
Autophagy is an evolutionarily conserved catabolic process that recycles nutrients upon starvation and maintains cellular energy homeostasis. Its acute regulation by nutrient-sensing signalling pathways is well described, but its longer-term transcriptional regulation is not. The nuclear receptors peroxisome proliferator-activated receptor-α (PPARα) and farnesoid X receptor (FXR) are activated in the fasted and fed liver, respectively. Here we show that both PPARα and FXR regulate hepatic autophagy in mice. Pharmacological activation of PPARα reverses the normal suppression of autophagy in the fed state, inducing autophagic lipid degradation, or lipophagy. This response is lost in PPARα knockout (Ppara(-/-), also known as Nr1c1(-/-)) mice, which are partially defective in the induction of autophagy by fasting. Pharmacological activation of the bile acid receptor FXR strongly suppresses the induction of autophagy in the fasting state, and this response is absent in FXR knockout (Fxr(-/-), also known as Nr1h4(-/-)) mice, which show a partial defect in suppression of hepatic autophagy in the fed state. PPARα and FXR compete for binding to shared sites in autophagic gene promoters, with opposite transcriptional outputs. These results reveal complementary, interlocking mechanisms for regulation of autophagy by nutrient status.
Figures





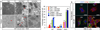

References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
- R01 DK043806/DK/NIDDK NIH HHS/United States
- U54HD-07495-39/HD/NICHD NIH HHS/United States
- S10RR027783-01A1/RR/NCRR NIH HHS/United States
- P30 CA125123/CA/NCI NIH HHS/United States
- P30 DK056338/DK/NIDDK NIH HHS/United States
- R01 DK049780/DK/NIDDK NIH HHS/United States
- S10 RR027783/RR/NCRR NIH HHS/United States
- DK43806/DK/NIDDK NIH HHS/United States
- P39CA125123-04/CA/NCI NIH HHS/United States
- U54 HD007495/HD/NICHD NIH HHS/United States
- R01 DK49780/DK/NIDDK NIH HHS/United States
- R37 DK043806/DK/NIDDK NIH HHS/United States
- P30 DK019525/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

