TP53 oncomorphic mutations predict resistance to platinum‑ and taxane‑based standard chemotherapy in patients diagnosed with advanced serous ovarian carcinoma
- PMID: 25385265
- PMCID: PMC4277253
- DOI: 10.3892/ijo.2014.2747
TP53 oncomorphic mutations predict resistance to platinum‑ and taxane‑based standard chemotherapy in patients diagnosed with advanced serous ovarian carcinoma
Abstract
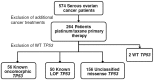
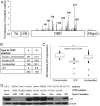
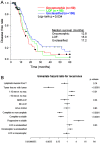
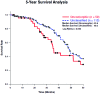
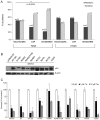
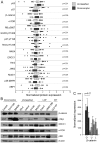
Individual mutations in the tumor suppressor TP53 alter p53 protein function. Some mutations create a non-functional protein, whereas others confer oncogenic activity, which we term 'oncomorphic'. Since mutations in TP53 occur in nearly all ovarian tumors, the objective of this study was to determine the relationship of oncomorphic TP53 mutations with patient outcomes in advanced serous ovarian cancer patients. Clinical and molecular data from 264 high-grade serous ovarian cancer patients uniformly treated with standard platinum- and taxane-based adjuvant chemotherapy were downloaded from The Cancer Genome Atlas (TCGA) portal. Additionally, patient samples were obtained from the University of Iowa and individual mutations were analyzed in ovarian cancer cell lines. Mutations in the TP53 were annotated and categorized as oncomorphic, loss of function (LOF), or unclassified. Associations between mutation types, chemoresistance, recurrence, and progression-free survival (PFS) were calculated. Oncomorphic TP53 mutations were present in 21.3% of ovarian cancers in the TCGA dataset. Patients with oncomorphic TP53 mutations demonstrated significantly worse PFS, a 60% higher risk of recurrence (HR=1.60, 95% confidence intervals 1.09, 2.33, p=0.015), and higher rates of platinum resistance (χ(2) test p=0.0024) when compared with single nucleotide mutations not categorized as oncomorphic. Furthermore, tumors containing oncomorphic TP53 mutations displayed unique protein expression profiles, and some mutations conferred increased clonogenic capacity in ovarian cancer cell models. Our study reveals that oncomorphic TP53 mutations are associated with worse patient outcome. These data suggest that future studies should take into consideration the functional consequences of TP53 mutations when determining treatment options.
Figures
Similar articles
-
TP53 mutations are common in all subtypes of epithelial ovarian cancer and occur concomitantly with KRAS mutations in the mucinous type.Exp Mol Pathol. 2013 Oct;95(2):235-41. doi: 10.1016/j.yexmp.2013.08.004. Epub 2013 Aug 18. Exp Mol Pathol. 2013. PMID: 23965232
-
Keratin 5 overexpression is associated with serous ovarian cancer recurrence and chemotherapy resistance.Oncotarget. 2017 Mar 14;8(11):17819-17832. doi: 10.18632/oncotarget.14867. Oncotarget. 2017. PMID: 28147318 Free PMC article.
-
TP53 mutations and the association with platinum resistance in high grade serous ovarian carcinoma.Gynecol Oncol. 2024 Jul;186:26-34. doi: 10.1016/j.ygyno.2024.03.023. Epub 2024 Mar 30. Gynecol Oncol. 2024. PMID: 38555766 Free PMC article.
-
The consequence of oncomorphic TP53 mutations in ovarian cancer.Int J Mol Sci. 2013 Sep 23;14(9):19257-75. doi: 10.3390/ijms140919257. Int J Mol Sci. 2013. PMID: 24065105 Free PMC article. Review.
-
Low Grade Serous Ovarian Carcinoma: from the molecular characterization to the best therapeutic strategy.Cancer Treat Rev. 2015 Feb;41(2):136-43. doi: 10.1016/j.ctrv.2014.12.003. Epub 2014 Dec 23. Cancer Treat Rev. 2015. PMID: 25573350 Review.
Cited by
-
Gain-of-Function Mutant TP53 R248Q Overexpressed in Epithelial Ovarian Carcinoma Alters AKT-Dependent Regulation of Intercellular Trafficking in Responses to EGFR/MDM2 Inhibitor.Int J Mol Sci. 2021 Aug 16;22(16):8784. doi: 10.3390/ijms22168784. Int J Mol Sci. 2021. PMID: 34445495 Free PMC article.
-
p53 mutation status is a primary determinant of placenta-specific protein 1 expression in serous ovarian cancers.Int J Oncol. 2017 May;50(5):1721-1728. doi: 10.3892/ijo.2017.3931. Epub 2017 Mar 23. Int J Oncol. 2017. PMID: 28339050 Free PMC article.
-
Insight updating of the molecular hallmarks in ovarian carcinoma.EJC Suppl. 2020 Aug 22;15:16-26. doi: 10.1016/j.ejcsup.2019.11.001. eCollection 2020 Aug. EJC Suppl. 2020. PMID: 33240439 Free PMC article.
-
Clinical Outcomes of TP53 Mutations in Cancers.Cold Spring Harb Perspect Med. 2016 Sep 1;6(9):a026294. doi: 10.1101/cshperspect.a026294. Cold Spring Harb Perspect Med. 2016. PMID: 27449973 Free PMC article. Review.
-
Landscape of genomic alterations in high-grade serous ovarian cancer from exceptional long- and short-term survivors.Genome Med. 2018 Oct 31;10(1):81. doi: 10.1186/s13073-018-0590-x. Genome Med. 2018. PMID: 30382883 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

