Next-generation sequencing in veterinary medicine: how can the massive amount of information arising from high-throughput technologies improve diagnosis, control, and management of infectious diseases?
- PMID: 25399113
- PMCID: PMC7123048
- DOI: 10.1007/978-1-4939-2004-4_30
Next-generation sequencing in veterinary medicine: how can the massive amount of information arising from high-throughput technologies improve diagnosis, control, and management of infectious diseases?
Abstract
The development of high-throughput molecular technologies and associated bioinformatics has dramatically changed the capacities of scientists to produce, handle, and analyze large amounts of genomic, transcriptomic, and proteomic data. A clear example of this step-change is represented by the amount of DNA sequence data that can be now produced using next-generation sequencing (NGS) platforms. Similarly, recent improvements in protein and peptide separation efficiencies and highly accurate mass spectrometry have promoted the identification and quantification of proteins in a given sample. These advancements in biotechnology have increasingly been applied to the study of animal infectious diseases and are beginning to revolutionize the way that biological and evolutionary processes can be studied at the molecular level. Studies have demonstrated the value of NGS technologies for molecular characterization, ranging from metagenomic characterization of unknown pathogens or microbial communities to molecular epidemiology and evolution of viral quasispecies. Moreover, high-throughput technologies now allow detailed studies of host-pathogen interactions at the level of their genomes (genomics), transcriptomes (transcriptomics), or proteomes (proteomics). Ultimately, the interaction between pathogen and host biological networks can be questioned by analytically integrating these levels (integrative OMICS and systems biology). The application of high-throughput biotechnology platforms in these fields and their typical low-cost per information content has revolutionized the resolution with which these processes can now be studied. The aim of this chapter is to provide a current and prospective view on the opportunities and challenges associated with the application of massive parallel sequencing technologies to veterinary medicine, with particular focus on applications that have a potential impact on disease control and management.
Figures


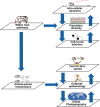
Similar articles
-
Novel technologies applied to the nucleotide sequencing and comparative sequence analysis of the genomes of infectious agents in veterinary medicine.Rev Sci Tech. 2016 Apr;35(1):25-42. doi: 10.20506/rst.35.1.2415. Rev Sci Tech. 2016. PMID: 27217166 Review.
-
High-throughput sequencing in veterinary infection biology and diagnostics.Rev Sci Tech. 2013 Dec;32(3):893-915. doi: 10.20506/rst.32.2.2206. Rev Sci Tech. 2013. PMID: 24761741
-
Proteomics in veterinary medicine: applications and trends in disease pathogenesis and diagnostics.Vet Pathol. 2014 Mar;51(2):351-62. doi: 10.1177/0300985813502819. Epub 2013 Sep 17. Vet Pathol. 2014. PMID: 24045891 Review.
-
Genomics and vaccine development.Rev Sci Tech. 2007 Apr;26(1):49-67. Rev Sci Tech. 2007. PMID: 17633293 Review.
-
The Global Genome Question: Microbes as the Key to Understanding Evolution and Ecology: This report is based on a colloquium, “The Global Genome Question: Microbes as the Key to Understanding Evolution and Ecology,” sponsored by the American Academy of Microbiology and held October 11-13, 2002, in Longboat Key, Florida.Washington (DC): American Society for Microbiology; 2004. Washington (DC): American Society for Microbiology; 2004. PMID: 33119236 Free Books & Documents. Review.
Cited by
-
Compartmentalized evolution of Bovine Viral Diarrhoea Virus type 2 in an immunotolerant persistently infected cow.Sci Rep. 2019 Oct 29;9(1):15460. doi: 10.1038/s41598-019-52023-w. Sci Rep. 2019. PMID: 31664116 Free PMC article.
-
Next-Generation Sequencing for the Detection of Microbial Agents in Avian Clinical Samples.Vet Sci. 2023 Dec 4;10(12):690. doi: 10.3390/vetsci10120690. Vet Sci. 2023. PMID: 38133241 Free PMC article. Review.
-
Microbiome Research as an Effective Driver of Success Stories in Agrifood Systems - A Selection of Case Studies.Front Microbiol. 2022 Jul 4;13:834622. doi: 10.3389/fmicb.2022.834622. eCollection 2022. Front Microbiol. 2022. PMID: 35903477 Free PMC article. Review.
-
Detection of Mycobacteria in Arabian camels and antimycobacterial potential of Moringa oleifera.Sci Rep. 2025 Apr 4;15(1):11590. doi: 10.1038/s41598-025-92402-0. Sci Rep. 2025. PMID: 40185780 Free PMC article.
-
Challenges and Solutions to Viral Diseases of Finfish in Marine Aquaculture.Pathogens. 2021 May 30;10(6):673. doi: 10.3390/pathogens10060673. Pathogens. 2021. PMID: 34070735 Free PMC article. Review.
References
-
- Mullis K, Faloona F, Scharf S, et al. Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harb Symp Quant Biol. 1986;51:263–273. - PubMed
-
- Bartlett JM, Stirling D. A short history of the polymerase chain reaction. Methods Mol Biol. 2003;226:3–6. - PubMed
-
- Glenn TC. Field guide to next-generation DNA sequencers. Mol Ecol Resour. 2011;11:759–769. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

