CCDC88B is a novel regulator of maturation and effector functions of T cells during pathological inflammation
- PMID: 25403443
- PMCID: PMC4267237
- DOI: 10.1084/jem.20140455
CCDC88B is a novel regulator of maturation and effector functions of T cells during pathological inflammation
Abstract
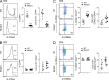
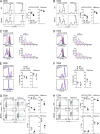
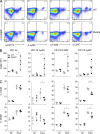
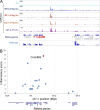
We used a genome-wide screen in mutagenized mice to identify genes which inactivation protects against lethal neuroinflammation during experimental cerebral malaria (ECM). We identified an ECM-protective mutation in coiled-coil domain containing protein 88b (Ccdc88b), a poorly annotated gene that is found expressed specifically in spleen, bone marrow, lymph nodes, and thymus. The CCDC88B protein is abundantly expressed in immune cells, including both CD4(+) and CD8(+) T lymphocytes, and in myeloid cells, and loss of CCDC88B protein expression has pleiotropic effects on T lymphocyte functions, including impaired maturation in vivo, significantly reduced activation, reduced cell division as well as impaired cytokine production (IFN-γ and TNF) in response to T cell receptor engagement, or to nonspecific stimuli in vitro, and during the course of P. berghei infection in vivo. This identifies CCDC88B as a novel and important regulator of T cell function. The human CCDC88B gene maps to the 11q13 locus that is associated with susceptibility to several inflammatory and auto-immune disorders. Our findings strongly suggest that CCDC88B is the morbid gene underlying the pleiotropic effect of the 11q13 locus on inflammation.
© 2014 Kennedy et al.
Figures




Comment in
-
A novel immune regulator links malaria and inflammatory bowel disease.J Exp Med. 2014 Dec 15;211(13):2482. doi: 10.1084/jem.21113insight1. J Exp Med. 2014. PMID: 25512579 Free PMC article. No abstract available.
Similar articles
-
CCDC88B is required for pathogenesis of inflammatory bowel disease.Nat Commun. 2017 Oct 13;8(1):932. doi: 10.1038/s41467-017-01381-y. Nat Commun. 2017. PMID: 29030607 Free PMC article.
-
CCDC88B is required for mobility and inflammatory functions of dendritic cells.J Leukoc Biol. 2020 Dec;108(6):1787-1802. doi: 10.1002/JLB.3A0420-386R. Epub 2020 Jun 1. J Leukoc Biol. 2020. PMID: 32480428
-
IRGM3 contributes to immunopathology and is required for differentiation of antigen-specific effector CD8+ T cells in experimental cerebral malaria.Infect Immun. 2015 Apr;83(4):1406-17. doi: 10.1128/IAI.02701-14. Epub 2015 Feb 2. Infect Immun. 2015. PMID: 25644000 Free PMC article.
-
The transcription factor T-bet regulates parasitemia and promotes pathogenesis during Plasmodium berghei ANKA murine malaria.J Immunol. 2013 Nov 1;191(9):4699-708. doi: 10.4049/jimmunol.1300396. Epub 2013 Sep 27. J Immunol. 2013. PMID: 24078698
-
THEMIS is required for pathogenesis of cerebral malaria and protection against pulmonary tuberculosis.Infect Immun. 2015 Feb;83(2):759-68. doi: 10.1128/IAI.02586-14. Epub 2014 Dec 1. Infect Immun. 2015. PMID: 25452553 Free PMC article.
Cited by
-
Blood and Intestine eQTLs from an Anti-TNF-Resistant Crohn's Disease Cohort Inform IBD Genetic Association Loci.Clin Transl Gastroenterol. 2016 Jun 23;7(6):e177. doi: 10.1038/ctg.2016.34. Clin Transl Gastroenterol. 2016. PMID: 27336838 Free PMC article.
-
CCDC88B interacts with RASAL3 and ARHGEF2 and regulates dendritic cell function in neuroinflammation and colitis.Commun Biol. 2024 Jan 10;7(1):77. doi: 10.1038/s42003-023-05751-9. Commun Biol. 2024. PMID: 38200184 Free PMC article.
-
GWAS identifies genetic loci, lifestyle factors and circulating biomarkers that are risk factors for sarcoidosis.Nat Commun. 2025 Mar 12;16(1):2481. doi: 10.1038/s41467-025-57829-z. Nat Commun. 2025. PMID: 40075078 Free PMC article.
-
Discovery of new candidate genes for rheumatoid arthritis through integration of genetic association data with expression pathway analysis.Arthritis Res Ther. 2017 Feb 2;19(1):19. doi: 10.1186/s13075-017-1220-5. Arthritis Res Ther. 2017. PMID: 28148290 Free PMC article.
-
Multi-ancestry and multi-trait genome-wide association meta-analyses inform clinical risk prediction for systemic lupus erythematosus.Nat Commun. 2023 Feb 7;14(1):668. doi: 10.1038/s41467-023-36306-5. Nat Commun. 2023. PMID: 36750564 Free PMC article.
References
-
- Beecham A.H., Patsopoulos N.A., Xifara D.K., Davis M.F., Kemppinen A., Cotsapas C., Shah T.S., Spencer C., Booth D., Goris A., et al. . International IBD Genetics Consortium (IIBDGC). 2013. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 45:1353–1360 10.1038/ng.2770 - DOI - PMC - PubMed
-
- Bongfen S.E., Rodrigue-Gervais I.-G., Berghout J., Torre S., Cingolani P., Wiltshire S.A., Leiva-Torres G.A., Letourneau L., Sladek R., Blanchette M., et al. . 2012. An N-ethyl-N-nitrosourea (ENU)-induced dominant negative mutation in the JAK3 kinase protects against cerebral malaria. PLoS ONE. 7:e31012 10.1371/journal.pone.0031012 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

