An enteric virus can replace the beneficial function of commensal bacteria
- PMID: 25409145
- PMCID: PMC4257755
- DOI: 10.1038/nature13960
An enteric virus can replace the beneficial function of commensal bacteria
Abstract
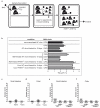
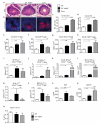
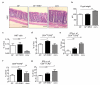
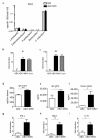
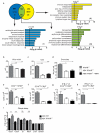
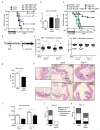
Intestinal microbial communities have profound effects on host physiology. Whereas the symbiotic contribution of commensal bacteria is well established, the role of eukaryotic viruses that are present in the gastrointestinal tract under homeostatic conditions is undefined. Here we demonstrate that a common enteric RNA virus can replace the beneficial function of commensal bacteria in the intestine. Murine norovirus (MNV) infection of germ-free or antibiotic-treated mice restored intestinal morphology and lymphocyte function without inducing overt inflammation and disease. The presence of MNV also suppressed an expansion of group 2 innate lymphoid cells observed in the absence of bacteria, and induced transcriptional changes in the intestine associated with immune development and type I interferon (IFN) signalling. Consistent with this observation, the IFN-α receptor was essential for the ability of MNV to compensate for bacterial depletion. Importantly, MNV infection offset the deleterious effect of treatment with antibiotics in models of intestinal injury and pathogenic bacterial infection. These data indicate that eukaryotic viruses have the capacity to support intestinal homeostasis and shape mucosal immunity, similarly to commensal bacteria.
Figures




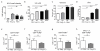

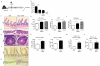

Comment in
-
Microbiology: a backup for bacteria.Nature. 2014 Dec 4;516(7529):42-3. doi: 10.1038/nature13938. Epub 2014 Nov 19. Nature. 2014. PMID: 25409141 No abstract available.
-
Microbiota: A viral understudy for commensal bacteria.Nat Rev Immunol. 2015 Jan;15(1):4. doi: 10.1038/nri3788. Epub 2014 Nov 28. Nat Rev Immunol. 2015. PMID: 25430963 No abstract available.
-
Gut microbiota: a 'friendly' gut virus?Nat Rev Gastroenterol Hepatol. 2015 Jan;12(1):6. doi: 10.1038/nrgastro.2014.220. Epub 2014 Dec 9. Nat Rev Gastroenterol Hepatol. 2015. PMID: 25488674 No abstract available.
References
-
- Donaldson EF, Lindesmith LC, Lobue AD, Baric RS. Norovirus pathogenesis: mechanisms of persistence and immune evasion in human populations. Immunol Rev. 2008;225:190–211. - PubMed
Supplemental references
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

