Exploring the distribution of genetic markers of pharmacogenomics relevance in Brazilian and Mexican populations
- PMID: 25419701
- PMCID: PMC4242606
- DOI: 10.1371/journal.pone.0112640
Exploring the distribution of genetic markers of pharmacogenomics relevance in Brazilian and Mexican populations
Erratum in
-
Correction: Exploring the distribution of genetic markers of pharmacogenomics relevance in Brazilian and Mexican populations.PLoS One. 2015 Mar 23;10(3):e0122161. doi: 10.1371/journal.pone.0122161. eCollection 2015. PLoS One. 2015. PMID: 25799410 Free PMC article. No abstract available.
Abstract
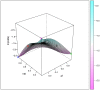
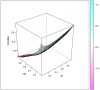
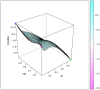
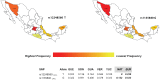
Studies of pharmacogenomics-related traits are increasingly being performed to identify loci that affect either drug response or susceptibility to adverse drug reactions. However, the effect of the polymorphisms can differ in magnitude or be absent depending on the population being assessed. We used the Affymetrix Drug Metabolizing Enzymes and Transporters (DMET) Plus array to characterize the distribution of polymorphisms of pharmacogenetics and pharmacogenomics (PGx) relevance in two samples from the most populous Latin American countries, Brazil and Mexico. The sample from Brazil included 268 individuals from the southeastern state of Rio de Janeiro, and was stratified into census categories. The sample from Mexico comprised 45 Native American Zapotecas and 224 self-identified Mestizo individuals from 5 states located in geographically distant regions in Mexico. We evaluated the admixture proportions in the Brazilian and Mexican samples using a panel of Ancestry Informative Markers extracted from the DMET array, which was validated with genome-wide data. A substantial variation in ancestral proportions across census categories in Brazil, and geographic regions in Mexico was identified. We evaluated the extent of genetic differentiation (measured as FST values) of the genetic markers of the DMET Plus array between the relevant parental populations. Although the average levels of genetic differentiation are low, there is a long tail of markers showing large frequency differences, including markers located in genes belonging to the Cytochrome P450, Solute Carrier (SLC) and UDP-glucuronyltransferase (UGT) families as well as other genes of PGx relevance such as ABCC8, ADH1A, CHST3, PON1, PPARD, PPARG, and VKORC1. We show how differences in admixture history may have an important impact in the distribution of allele and genotype frequencies at the population level.
Conflict of interest statement
Figures

Similar articles
-
Genotype frequencies of VKORC1 and CYP2C9 in native and Mestizo populations from Mexico, potential impact for coumarin dosing.Gene. 2015 Mar 10;558(2):235-40. doi: 10.1016/j.gene.2014.12.068. Epub 2015 Jan 2. Gene. 2015. PMID: 25560189
-
Distribution and linkage disequilibrium of the enhancer SNP rs5758550 among Latin American populations: influence of continental ancestry.Pharmacogenet Genomics. 2020 Jun;30(4):67-72. doi: 10.1097/FPC.0000000000000398. Pharmacogenet Genomics. 2020. PMID: 32187157
-
Population Diversity in Pharmacogenetics: A Latin American Perspective.Adv Pharmacol. 2018;83:133-154. doi: 10.1016/bs.apha.2018.02.001. Epub 2018 Mar 19. Adv Pharmacol. 2018. PMID: 29801573 Review.
-
Pharmacogenetics of drug metabolizing enzymes in Brazilian populations.Drug Metabol Drug Interact. 2014;29(3):153-77. doi: 10.1515/dmdi-2013-0067. Drug Metabol Drug Interact. 2014. PMID: 24744305 Review.
-
Exploring Variation in Known Pharmacogenetic Variants and its Association with Drug Response in Different Mexican Populations.Pharm Res. 2016 Nov;33(11):2644-52. doi: 10.1007/s11095-016-1990-5. Epub 2016 Jul 7. Pharm Res. 2016. PMID: 27387170
Cited by
-
A European Spectrum of Pharmacogenomic Biomarkers: Implications for Clinical Pharmacogenomics.PLoS One. 2016 Sep 16;11(9):e0162866. doi: 10.1371/journal.pone.0162866. eCollection 2016. PLoS One. 2016. PMID: 27636550 Free PMC article.
-
Population Pharmacogenomics for Precision Public Health in Colombia.Front Genet. 2019 Mar 22;10:241. doi: 10.3389/fgene.2019.00241. eCollection 2019. Front Genet. 2019. PMID: 30967898 Free PMC article.
-
Identification of ancestry proportions in admixed groups across the Americas using clinical pharmacogenomic SNP panels.Sci Rep. 2021 Jan 13;11(1):1007. doi: 10.1038/s41598-020-80389-9. Sci Rep. 2021. PMID: 33441860 Free PMC article.
-
Population structure and pharmacogenomic risk stratification in the United States.BMC Biol. 2020 Oct 13;18(1):140. doi: 10.1186/s12915-020-00875-4. BMC Biol. 2020. PMID: 33050895 Free PMC article.
-
Distribution of a novel CYP2C haplotype in Native American populations.Front Genet. 2023 Mar 21;14:1114742. doi: 10.3389/fgene.2023.1114742. eCollection 2023. Front Genet. 2023. PMID: 37025454 Free PMC article.
References
-
- The Single Nucleotide Polymorphism database website. Available: http://www.ncbi.nlm.nih.gov/snp/?term=9923231. Accessed 2013 December 17.
-
- Rede Nacional de Farmacogenética website. Available: http://www.refargen.org.br/article.php3?id_article=50. Accessed 2014 January 14.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

