The first myriapod genome sequence reveals conservative arthropod gene content and genome organisation in the centipede Strigamia maritima
- PMID: 25423365
- PMCID: PMC4244043
- DOI: 10.1371/journal.pbio.1002005
The first myriapod genome sequence reveals conservative arthropod gene content and genome organisation in the centipede Strigamia maritima
Abstract
Myriapods (e.g., centipedes and millipedes) display a simple homonomous body plan relative to other arthropods. All members of the class are terrestrial, but they attained terrestriality independently of insects. Myriapoda is the only arthropod class not represented by a sequenced genome. We present an analysis of the genome of the centipede Strigamia maritima. It retains a compact genome that has undergone less gene loss and shuffling than previously sequenced arthropods, and many orthologues of genes conserved from the bilaterian ancestor that have been lost in insects. Our analysis locates many genes in conserved macro-synteny contexts, and many small-scale examples of gene clustering. We describe several examples where S. maritima shows different solutions from insects to similar problems. The insect olfactory receptor gene family is absent from S. maritima, and olfaction in air is likely effected by expansion of other receptor gene families. For some genes S. maritima has evolved paralogues to generate coding sequence diversity, where insects use alternate splicing. This is most striking for the Dscam gene, which in Drosophila generates more than 100,000 alternate splice forms, but in S. maritima is encoded by over 100 paralogues. We see an intriguing linkage between the absence of any known photosensory proteins in a blind organism and the additional absence of canonical circadian clock genes. The phylogenetic position of myriapods allows us to identify where in arthropod phylogeny several particular molecular mechanisms and traits emerged. For example, we conclude that juvenile hormone signalling evolved with the emergence of the exoskeleton in the arthropods and that RR-1 containing cuticle proteins evolved in the lineage leading to Mandibulata. We also identify when various gene expansions and losses occurred. The genome of S. maritima offers us a unique glimpse into the ancestral arthropod genome, while also displaying many adaptations to its specific life history.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures






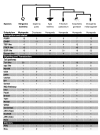



Comment in
-
What goes "99-thump?".PLoS Biol. 2014 Nov 25;12(11):e1002006. doi: 10.1371/journal.pbio.1002006. eCollection 2014 Nov. PLoS Biol. 2014. PMID: 25422891 Free PMC article. No abstract available.
-
Genome evolution: groping in the soil interstices.Curr Biol. 2015 Mar 2;25(5):R194-6. doi: 10.1016/j.cub.2015.01.018. Curr Biol. 2015. PMID: 25734267
References
-
- Arthropod Genomes Consortium (2014) List of sequenced arthropod genomes. Available: http://arthropodgenomes.org/wiki/Sequenced_genomes.
-
- Edgecombe GD (2011) Phylogenetic relationships of Myriapoda. Minelli A, editor. The Myriapoda. Leiden: Brill. pp. 1–20.
-
- Giribet G, Edgecombe GD, Wheeler WC (2001) Arthropod phylogeny based on eight molecular loci and morphology. Nature 157–160. - PubMed
-
- Rota-Stabelli O, Telford MJ (2008) A multi criterion approach for the selection of optimal outgroups in phylogeny: recovering some support for Mandibulata over Myriochelata using mitogenomics. Mol Phylogenet Evol 48: 103–111. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

