Structural characterization of the late competence protein ComFB from Bacillus subtilis
- PMID: 25423369
- PMCID: PMC4381287
- DOI: 10.1042/BSR20140174
Structural characterization of the late competence protein ComFB from Bacillus subtilis
Abstract
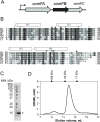
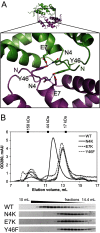
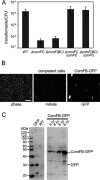
Many bacteria take up DNA from their environment as part of the process of natural transformation. DNA uptake allows microorganisms to gain genetic diversity and can lead to the spread of antibiotic resistance or virulence genes within a microbial population. Development of genetic competence (Com) in Bacillus subtilis is a highly regulated process that culminates in expression of several late competence genes and formation of the DNA uptake apparatus. The late competence operon comF encodes a small protein of unknown function, ComFB. To gain insight into the function of ComFB, we determined its 3D structure via X-ray crystallography. ComFB is a dimer and each subunit consists of four α-helices connected by short loops and one extended β-strand-like stretch. Each subunit contains one zinc-binding site formed by four cysteines, which are unusually spaced in the primary sequence. Using structure- and bioinformatics-guided substitutions we analyzed the inter-subunit interface of the ComFB dimer. Based on these analyses, we conclude that ComFB is an obligate dimer. We also characterized ComFB in vivo and found that this protein is produced in competent cells and is localized to the cytosol. Consistent with previous reports, we showed that deletion of ComFB does not affect DNA uptake function. Combining our results, we conclude that ComFB is unlikely to be a part of the DNA uptake machinery under tested conditions and instead may have a regulatory function.
Figures
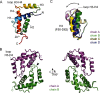


Similar articles
-
A Conserved Metal Binding Motif in the Bacillus subtilis Competence Protein ComFA Enhances Transformation.J Bacteriol. 2017 Jul 11;199(15):e00272-17. doi: 10.1128/JB.00272-17. Print 2017 Aug 1. J Bacteriol. 2017. PMID: 28559293 Free PMC article.
-
Fluorescently Labeled DNA Interacts with Competence and Recombination Proteins and Is Integrated and Expressed Following Natural Transformation of Bacillus subtilis.mBio. 2018 Sep 25;9(5):e01161-18. doi: 10.1128/mBio.01161-18. mBio. 2018. PMID: 30254116 Free PMC article.
-
Structure and organisation of SinR, the master regulator of biofilm formation in Bacillus subtilis.J Mol Biol. 2011 Aug 19;411(3):597-613. doi: 10.1016/j.jmb.2011.06.004. Epub 2011 Jun 25. J Mol Biol. 2011. PMID: 21708175 Free PMC article.
-
Genetic competence in Bacillus subtilis.Microbiol Rev. 1991 Sep;55(3):395-424. doi: 10.1128/mr.55.3.395-424.1991. Microbiol Rev. 1991. PMID: 1943994 Free PMC article. Review.
-
Ubiquitous late competence genes in Bacillus species indicate the presence of functional DNA uptake machineries.Environ Microbiol. 2009 Aug;11(8):1911-22. doi: 10.1111/j.1462-2920.2009.01937.x. Epub 2009 May 14. Environ Microbiol. 2009. PMID: 19453701 Review.
Cited by
-
A c-di-GMP binding effector controls cell size in a cyanobacterium.Proc Natl Acad Sci U S A. 2023 Mar 28;120(13):e2221874120. doi: 10.1073/pnas.2221874120. Epub 2023 Mar 22. Proc Natl Acad Sci U S A. 2023. PMID: 36947515 Free PMC article.
-
ComFB, a new widespread family of c-di-NMP receptor proteins.bioRxiv [Preprint]. 2024 Nov 10:2024.11.10.622515. doi: 10.1101/2024.11.10.622515. bioRxiv. 2024. PMID: 39574629 Free PMC article. Preprint.
-
c-di-GMP-Dependent Regulation of Motility by comFB and comFC.bioRxiv [Preprint]. 2025 Jul 11:2025.07.11.664319. doi: 10.1101/2025.07.11.664319. bioRxiv. 2025. PMID: 40672238 Free PMC article. Preprint.
-
Proteins identified through predictive metagenomics as potential biomarkers for the detection of microbiologically influenced corrosion.J Ind Microbiol Biotechnol. 2022 Jan 20;49(1):kuab068. doi: 10.1093/jimb/kuab068. J Ind Microbiol Biotechnol. 2022. PMID: 34543407 Free PMC article.
-
ComEB protein is dispensable for the transformation but must be translated for the optimal synthesis of comEC.Mol Microbiol. 2021 Jul;116(1):71-79. doi: 10.1111/mmi.14690. Epub 2021 Feb 8. Mol Microbiol. 2021. PMID: 33527432 Free PMC article.
References
-
- U.S. Department of Health and Human Services. 2013. Antibiotic Resistance Threats in the United States, Centers for Disease Control and Prevention, Atlanta. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

