The structure and regulation of human muscle α-actinin
- PMID: 25433700
- PMCID: PMC4259493
- DOI: 10.1016/j.cell.2014.10.056
The structure and regulation of human muscle α-actinin
Abstract
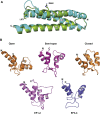
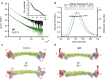
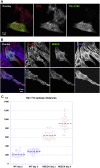
The spectrin superfamily of proteins plays key roles in assembling the actin cytoskeleton in various cell types, crosslinks actin filaments, and acts as scaffolds for the assembly of large protein complexes involved in structural integrity and mechanosensation, as well as cell signaling. α-actinins in particular are the major actin crosslinkers in muscle Z-disks, focal adhesions, and actin stress fibers. We report a complete high-resolution structure of the 200 kDa α-actinin-2 dimer from striated muscle and explore its functional implications on the biochemical and cellular level. The structure provides insight into the phosphoinositide-based mechanism controlling its interaction with sarcomeric proteins such as titin, lays a foundation for studying the impact of pathogenic mutations at molecular resolution, and is likely to be broadly relevant for the regulation of spectrin-like proteins.
Copyright © 2014 The Authors. Published by Elsevier Inc. All rights reserved.
Figures












References
-
- Atkinson R.A., Joseph C., Kelly G., Muskett F.W., Frenkiel T.A., Nietlispach D., Pastore A. Ca2+-independent binding of an EF-hand domain to a novel motif in the α-actinin-titin complex. Nat. Struct. Biol. 2001;8:853–857. - PubMed
-
- Bennett V., Healy J. Organizing the fluid membrane bilayer: diseases linked to spectrin and ankyrin. Trends Mol. Med. 2008;14:28–36. - PubMed
-
- Bernadó P., Mylonas E., Petoukhov M.V., Blackledge M., Svergun D.I. Structural characterization of flexible proteins using small-angle X-ray scattering. J. Am. Chem. Soc. 2007;129:5656–5664. - PubMed
Supplemental References
-
- Berendsen H.J.C., Postma J.P.M., van Gunsteren W.F., Hermans J. In: Intermolecular Forces. Pullman B., editor. Reidel; Dordrecht: 1981. Interaction models for water in relation to protein hydration; pp. 331–342.
-
- Berendsen H.J.C., Postma J.P.M., van Gunsteren W.F., DiNola A., Haak J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984;81:3684–3690.
-
- Blanchet C.E., Zozulya A.V., Kikhney A.G., Franke D., Konarev P.V., Shang W.F., Klaering R., Robrahn B., Hermes C., Cipriani F. Instrumental setup for high-throughput small- and wide-angle solution scattering at the X33 beamline of EMBL Hamburg. J. Appl. Crystallogr. 2012;45:489–495.
-
- Celniker G., Nimrod G., Ashkenazy H., Glaser F., Martz E., Mayrose I., Pupko T., Ben-Tal N. ConSurf: using evolutionary data to raise testable hypotheses about protein function. Isr. J. Chem. 2013;53:199–206.
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

