An ontology for microbial phenotypes
- PMID: 25433798
- PMCID: PMC4287307
- DOI: 10.1186/s12866-014-0294-3
An ontology for microbial phenotypes
Abstract
Background: Phenotypic data are routinely used to elucidate gene function in organisms amenable to genetic manipulation. However, previous to this work, there was no generalizable system in place for the structured storage and retrieval of phenotypic information for bacteria.
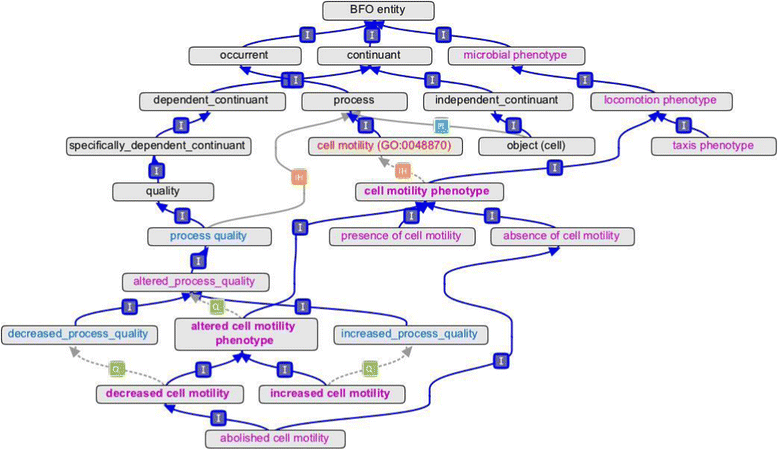
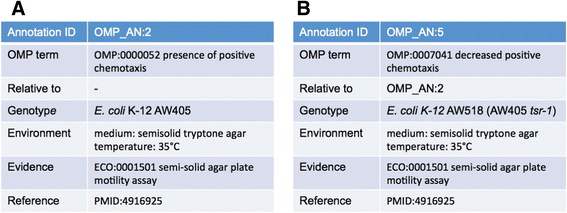
Results: The Ontology of Microbial Phenotypes (OMP) has been created to standardize the capture of such phenotypic information from microbes. OMP has been built on the foundations of the Basic Formal Ontology and the Phenotype and Trait Ontology. Terms have logical definitions that can facilitate computational searching of phenotypes and their associated genes. OMP can be accessed via a wiki page as well as downloaded from SourceForge. Initial annotations with OMP are being made for Escherichia coli using a wiki-based annotation capture system. New OMP terms are being concurrently developed as annotation proceeds.
Conclusions: We anticipate that diverse groups studying microbial genetics and associated phenotypes will employ OMP for standardizing microbial phenotype annotation, much as the Gene Ontology has standardized gene product annotation. The resulting OMP resource and associated annotations will facilitate prediction of phenotypes for unknown genes and result in new experimental characterization of phenotypes and functions.
Figures


References
-
- Holt JG. Bergey’s Manual of Determinative Microbiology. 9. Hagerstown, MD: Lippincott Williams & Wilkins; 1994.
-
- Smith B, Ashburner M, Rosse C, Bard J, Bug W, Ceusters W, Goldberg LJ, Eilbeck K, Ireland A, Mungall CJ, Leontis N, Rocca-Serra P, Ruttenberg A, Sansone SA, Scheuermann RH, Shah N, Whetzel PL, Lewis S. The OBO Foundry: coordinated evolution of ontologies to support biomedical data integration. Nat Biotechnol. 2007;25(11):1251–1255. doi: 10.1038/nbt1346. - DOI - PMC - PubMed
-
- Blom EJ, Breitling R, Hofstede KJ, Roerdink JB, Van Hijum SA, Kuipers OP. Prosecutor: parameter-free inference of gene function for prokaryotes using DNA microarray data, genomic context and multiple gene annotation sources. BMC Genomics. 2008;9:495. doi: 10.1186/1471-2164-9-495. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

