Comprehensive chronic lymphocytic leukemia diagnostics by combined multiplex ligation dependent probe amplification (MLPA) and interphase fluorescence in situ hybridization (iFISH)
- PMID: 25435911
- PMCID: PMC4247644
- DOI: 10.1186/s13039-014-0079-2
Comprehensive chronic lymphocytic leukemia diagnostics by combined multiplex ligation dependent probe amplification (MLPA) and interphase fluorescence in situ hybridization (iFISH)
Abstract
Background: Banding-karyotyping and metaphase-directed-fluorescence-in-situhybridization (FISH) may be hampered by low mitotic index in leukemia. Interphase FISH (iFISH) is a way out here, however, testing many probes at the same time is protracted and expensive. Here multiplex-ligation-dependent-probe-amplification (MLPA) was used retrospectively in chronic lymphocytic leukemia (CLL) samples initially studied by banding cytogenetics and iFISH. Detection rates of iFISH and MLPA were compared and thus a cost-efficient scheme for routine diagnostics is proposed.
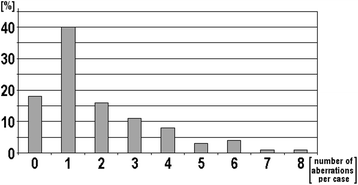
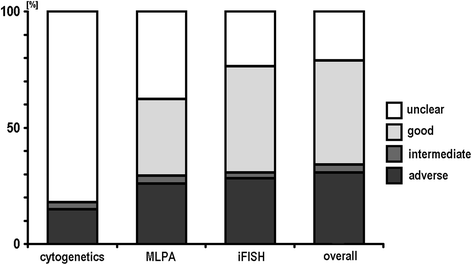
Results: Banding cytogenetics was done successfully in 67/85 samples. DNA was extracted from all 85 CLL samples. A commercially available MLPA probe set directed against 37 loci prone to be affected in hematological malignancies was applied. Besides, routine iFISH was done by commercially available probes for following regions: 11q22.3, 12p11.2-q11.1, 13q14.3, 13q34, 14q32.33 and 17p13.1. MLPA results were substantiated by iFISH using corresponding locus-specific probes. Aberrations were detected in 67 of 85 samples (~79%) applying banding cytogenetics, iFISH and MLPA. A maximum of 8 aberrations was detected per sample; however, one aberration per sample was found most frequently. Overall 163 aberrations were identified. 15 of those (~9%) were exclusively detected by banding cytogenetics, 95 were found by MLPA (~58%) and 100 (~61%) by routine iFISH. MLPA was not able to distinguish reliably between mono- and biallelic del(13)(q14.3q14.3), which could be easily identified as well as quantified by routine iFISH. Also iFISH was superior to MLPA in samples with low tumor cell load. On the other hand MLPA detected additional aberrations in 22 samples, two of them being without any findings after routine iFISH.
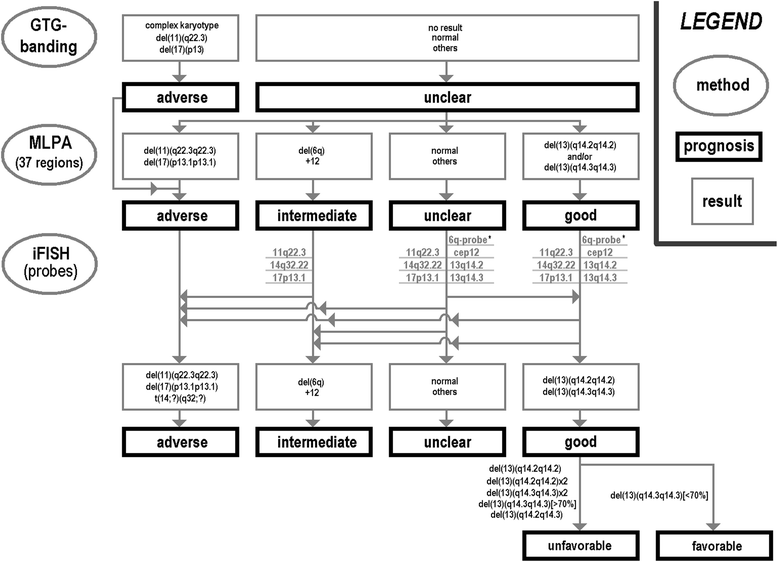
Conclusions: Both MLPA and routine iFISH have comparable detection rates for aberrations being typically present in CLL. As MLPA can detect also rare chromosomal aberrations it should be used as an initial test if routine cytogenetics is not possible or non-informative. Still iFISH should be used additionally to distinguish mono- from biallelic deletions and also to determine rate of mosaicism for 13q14.2 to 13q14.3. In case MLPA is negative the corresponding CLL samples should be tested at least by iFISH using the standard probe set to.
Keywords: Chromosomal aberrations; Chronic lymphocytic leukemia (CLL); Fluorescence in situ hybridization (FISH); Multiplex ligation-dependent probe amplification (MLPA).
Figures
Similar articles
-
Evaluation of MLPA as a comprehensive molecular cytogenetic tool to detect cytogenetic markers of chronic lymphocytic leukemia in Egyptian patients.J Genet Eng Biotechnol. 2021 Jun 28;19(1):98. doi: 10.1186/s43141-021-00198-z. J Genet Eng Biotechnol. 2021. PMID: 34181122 Free PMC article.
-
Interphase Molecular Cytogenetic Detection Rates of Chronic Lymphocytic Leukemia-Specific Aberrations Are Higher in Cultivated Cells Than in Blood or Bone Marrow Smears.J Histochem Cytochem. 2016 Aug;64(8):495-501. doi: 10.1369/0022155416655086. Epub 2016 Jun 17. J Histochem Cytochem. 2016. PMID: 27315825 Free PMC article.
-
High rates of submicroscopic aberrations in karyotypically normal acute lymphoblastic leukemia.Mol Cytogenet. 2015 Jun 30;8:45. doi: 10.1186/s13039-015-0153-4. eCollection 2015. Mol Cytogenet. 2015. PMID: 26136832 Free PMC article.
-
Genetic features of B-cell chronic lymphocytic leukemia.Rev Clin Exp Hematol. 2000 Mar;4(1):48-72. doi: 10.1046/j.1468-0734.2000.00003.x. Rev Clin Exp Hematol. 2000. PMID: 11486330 Review.
-
Cytogenetic and molecular cytogenetic analysis of B cell chronic lymphocytic leukemia: specific chromosome aberrations identify prognostic subgroups of patients and point to loci of candidate genes.Leukemia. 1997 Apr;11 Suppl 2:S19-24. Leukemia. 1997. PMID: 9178833 Review.
Cited by
-
Screening of the copy number increase of AKT in lung carcinoma by custom-designed MLPA.Int J Clin Exp Pathol. 2019 Sep 1;12(9):3344-3356. eCollection 2019. Int J Clin Exp Pathol. 2019. PMID: 31934177 Free PMC article.
-
BIRC3 alterations in chronic and B-cell acute lymphocytic leukemia patients.Oncol Lett. 2016 May;11(5):3240-3246. doi: 10.3892/ol.2016.4388. Epub 2016 Mar 29. Oncol Lett. 2016. PMID: 27123097 Free PMC article.
-
Evaluation of MLPA as a comprehensive molecular cytogenetic tool to detect cytogenetic markers of chronic lymphocytic leukemia in Egyptian patients.J Genet Eng Biotechnol. 2021 Jun 28;19(1):98. doi: 10.1186/s43141-021-00198-z. J Genet Eng Biotechnol. 2021. PMID: 34181122 Free PMC article.
-
Copy Number Variations and Gene Mutations Identified by Multiplex Ligation-Dependent Probe Amplification in Romanian Chronic Lymphocytic Leukemia Patients.J Pers Med. 2023 Aug 9;13(8):1239. doi: 10.3390/jpm13081239. J Pers Med. 2023. PMID: 37623489 Free PMC article.
-
Immunoglobulin gene translocations in chronic lymphocytic leukemia: A report of 35 patients and review of the literature.Mol Clin Oncol. 2016 May;4(5):682-694. doi: 10.3892/mco.2016.793. Epub 2016 Feb 26. Mol Clin Oncol. 2016. PMID: 27123263 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

