Evaluation of drug-drug interactions for oncology therapies: in vitro-in vivo extrapolation model-based risk assessment
- PMID: 25443889
- PMCID: PMC4456127
- DOI: 10.1111/bcp.12563
Evaluation of drug-drug interactions for oncology therapies: in vitro-in vivo extrapolation model-based risk assessment
Abstract
Aims: Understanding drug-drug interactions (DDI) is a critical part of the drug development process as polypharmacy has become commonplace in many therapeutic areas including the cancer patient population. The objectives of this study were to investigate cytochrome P450 (CYP)-mediated DDI profiles available for therapies used in the oncology setting and evaluate how models based on in vitro-in vivo extrapolation performed in predicting CYP-mediated DDI risk.
Methods: A dataset of 125 oncology therapies was collated using drug label and approval history information, incorporating in vitro and clinical PK data. The predictive accuracy of the basic and net effect mechanistic static models was assessed using this oncology drug dataset, for both victim and perpetrator potential of CYP3A-mediated DDI.
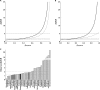
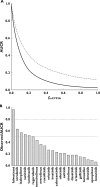
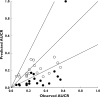
Results: The incidence of CYP3A-mediated interaction potential was 47%, 22% and 11% for substrates, inhibitors and inducers, respectively. The basic models for precipitants gave conservative predictions with no false negatives, whilst the mechanistic static models provided reasonable quantitative predictions (2.3-3-fold error). Further analysis revealed that incorporating DDI at the level of the intestine was in most cases over-predicting interaction magnitude due to overestimates of the rate and extent of oral absorption of the precipitant. Quantifying victim DDI potential was also demonstrated using fmCYP3A estimates from ketoconazole clinical DDI studies to predict the magnitude of interaction on co-administration with the CYP3A inducer, rifampicin (1.6-3.3 fold error).
Conclusions: This work illustrates the utility and limitations of current DDI risk assessment approaches applied to a range of contemporary anti-cancer agents, and discusses the implications for therapeutic combination strategies.
Keywords: CYP induction; CYP inhibition; CYP3A; anti-cancer therapy; drug interactions; pharmacokinetics.
© 2014 The British Pharmacological Society.
Figures

Similar articles
-
A mechanistic physiologically based pharmacokinetic-enzyme turnover model involving both intestine and liver to predict CYP3A induction-mediated drug-drug interactions.J Pharm Sci. 2013 Aug;102(8):2819-36. doi: 10.1002/jps.23613. Epub 2013 Jun 11. J Pharm Sci. 2013. PMID: 23760985
-
Application of Static Models to Predict Midazolam Clinical Interactions in the Presence of Single or Multiple Hepatitis C Virus Drugs.Drug Metab Dispos. 2016 Aug;44(8):1372-80. doi: 10.1124/dmd.116.070409. Epub 2016 May 25. Drug Metab Dispos. 2016. PMID: 27226352
-
Physiologically based pharmacokinetic modeling and simulation to predict drug-drug interactions of ivosidenib with CYP3A perpetrators in patients with acute myeloid leukemia.Cancer Chemother Pharmacol. 2020 Nov;86(5):619-632. doi: 10.1007/s00280-020-04148-3. Epub 2020 Sep 25. Cancer Chemother Pharmacol. 2020. PMID: 32978634
-
Alternatives to rifampicin: A review and perspectives on the choice of strong CYP3A inducers for clinical drug-drug interaction studies.Clin Transl Sci. 2022 Sep;15(9):2075-2095. doi: 10.1111/cts.13357. Epub 2022 Jul 25. Clin Transl Sci. 2022. PMID: 35722783 Free PMC article. Review.
-
Pharmacokinetic Interaction of Rifampicin with Oral Versus Intravenous Anticancer Drugs: Challenges, Dilemmas and Paradoxical Effects Due to Multiple Mechanisms.Drugs R D. 2016 Jun;16(2):141-8. doi: 10.1007/s40268-016-0133-0. Drugs R D. 2016. PMID: 27098526 Free PMC article. Review.
Cited by
-
Machine learning-based quantitative prediction of drug exposure in drug-drug interactions using drug label information.NPJ Digit Med. 2022 Jul 11;5(1):88. doi: 10.1038/s41746-022-00639-0. NPJ Digit Med. 2022. PMID: 35817846 Free PMC article.
-
Safety and Tolerability of Epidermal Growth Factor Receptor (EGFR) Tyrosine Kinase Inhibitors in Oncology.Drug Saf. 2019 Feb;42(2):181-198. doi: 10.1007/s40264-018-0772-x. Drug Saf. 2019. PMID: 30649743 Review.
-
Pharmacogenomic Characterization and Isobologram Analysis of the Combination of Ascorbic Acid and Curcumin-Two Main Metabolites of Curcuma longa-in Cancer Cells.Front Pharmacol. 2017 Feb 2;8:38. doi: 10.3389/fphar.2017.00038. eCollection 2017. Front Pharmacol. 2017. PMID: 28210221 Free PMC article.
-
Effect of fluconazole on the pharmacokinetics of a single dose of fedratinib in healthy adults.Cancer Chemother Pharmacol. 2022 Oct;90(4):325-334. doi: 10.1007/s00280-022-04464-w. Epub 2022 Aug 24. Cancer Chemother Pharmacol. 2022. PMID: 36001108 Free PMC article. Clinical Trial.
-
Evaluation of BCRP-Related DDIs Between Methotrexate and Cyclosporin A Using Physiologically Based Pharmacokinetic Modelling.Drugs R D. 2025 Mar;25(1):1-17. doi: 10.1007/s40268-024-00495-1. Epub 2024 Dec 24. Drugs R D. 2025. PMID: 39715910 Free PMC article.
References
-
- Jansman FGA, Reyners AKL, van Roon EN, Smorenburg CH, Helgason HH, Le Comte M, Wensveen BM, van den Tweel AMA, de Blois M, Kwee W, Kerremans AL, Brouwers JRBJ. Consensus-based evaluation of clinical significance and management of anticancer drug interactions. Clin Ther. 2011;33:305–14. - PubMed
-
- Kohler GI, Bode-Boger SM, Busse R, Hoopmann M, Welte T, Boger RH. Drug–drug interactions in medical patients: effects of in-hospital treatment and relation to multiple drug use. Int J Clin Pharmacol Ther. 2000;38:504–513. - PubMed
-
- Buajordet I, Ebbesen J, Erikssen J, Brors O, Hilberg T. Fatal adverse drug events: the paradox of drug treatment. J Intern Med. 2001;250:327–41. - PubMed
-
- Menzies AM, Long GV. Dabrafenib and trametinib, alone and in combination for BRAF-mutant metastatic melanoma. Clin Cancer Res. 2014;20:2035–43. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

