Integrative analysis of T cell motility from multi-channel microscopy data using TIAM
- PMID: 25445324
- PMCID: PMC4323926
- DOI: 10.1016/j.jim.2014.11.004
Integrative analysis of T cell motility from multi-channel microscopy data using TIAM
Abstract
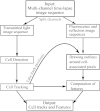
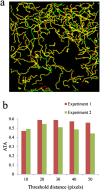
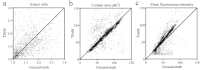
Integrative analytical approaches are needed to study and understand T cell motility as it is a highly coordinated and complex process. Several computational algorithms and tools are available to track motile cells in time-lapse microscopy images. In contrast, there has only been limited effort towards the development of tools that take advantage of multi-channel microscopy data and facilitate integrative analysis of cell-motility. We have implemented algorithms for detecting, tracking, and analyzing cell motility from multi-channel time-lapse microscopy data. We have integrated these into a MATLAB-based toolset we call TIAM (Tool for Integrative Analysis of Motility). The cells are detected by a hybrid approach involving edge detection and Hough transforms from transmitted light images. Cells are tracked using a modified nearest-neighbor association followed by an optimization routine to join shorter segments. Cell positions are used to perform local segmentation for extracting features from transmitted light, reflection and fluorescence channels and associating them with cells and cell-tracks to facilitate integrative analysis. We found that TIAM accurately captures the motility behavior of T cells and performed better than DYNAMIK, Icy, Imaris, and Volocity in detecting and tracking motile T cells. Extraction of cell-associated features from reflection and fluorescence channels was also accurate with less than 10% median error in measurements. Finally, we obtained novel insights into T cell motility that were critically dependent on the unique capabilities of TIAM. We found that 1) the CD45RO subset of human CD8 T cells moved faster and exhibited an increased propensity to attach to the substratum during CCL21-driven chemokinesis when compared to the CD45RA subset; and 2) attachment area and arrest coefficient during antigen-induced motility of the CD45A subset is correlated with surface density of integrin LFA1 at the contact.
Keywords: Integrative analysis; Multi-channel microscopy; T cell motility; Tracking.
Copyright © 2014 Elsevier B.V. All rights reserved.
Figures


Similar articles
-
An analytical tool that quantifies cellular morphology changes from three-dimensional fluorescence images.J Vis Exp. 2012 Aug 31;(66):e4233. doi: 10.3791/4233. J Vis Exp. 2012. PMID: 22951512 Free PMC article.
-
Visualization and correction of automated segmentation, tracking and lineaging from 5-D stem cell image sequences.BMC Bioinformatics. 2014 Oct 3;15(1):328. doi: 10.1186/1471-2105-15-328. BMC Bioinformatics. 2014. PMID: 25281197 Free PMC article.
-
Cell motility dynamics: a novel segmentation algorithm to quantify multi-cellular bright field microscopy images.PLoS One. 2011;6(11):e27593. doi: 10.1371/journal.pone.0027593. Epub 2011 Nov 9. PLoS One. 2011. PMID: 22096600 Free PMC article.
-
Computer-assisted systems for the analysis of amoeboid cell motility.Methods Mol Biol. 2001;161:45-58. doi: 10.1385/1-59259-051-9:045. Methods Mol Biol. 2001. PMID: 11190516 Review. No abstract available.
-
Untangling cell tracks: Quantifying cell migration by time lapse image data analysis.Cytometry A. 2018 Mar;93(3):357-370. doi: 10.1002/cyto.a.23249. Epub 2017 Oct 4. Cytometry A. 2018. PMID: 28976646 Review.
Cited by
-
In vitro machine learning-based CAR T immunological synapse quality measurements correlate with patient clinical outcomes.PLoS Comput Biol. 2022 Mar 18;18(3):e1009883. doi: 10.1371/journal.pcbi.1009883. eCollection 2022 Mar. PLoS Comput Biol. 2022. PMID: 35303007 Free PMC article. Clinical Trial.
-
T Lymphocyte Migration: An Action Movie Starring the Actin and Associated Actors.Front Immunol. 2015 Nov 18;6:586. doi: 10.3389/fimmu.2015.00586. eCollection 2015. Front Immunol. 2015. PMID: 26635800 Free PMC article. Review.
-
A tissue-like platform for studying engineered quiescent human T-cells' interactions with dendritic cells.Elife. 2019 Sep 25;8:e48221. doi: 10.7554/eLife.48221. Elife. 2019. PMID: 31552826 Free PMC article.
-
Interpreting T-cell search "strategies" in the light of evolution under constraints.PLoS Comput Biol. 2023 Feb 27;19(2):e1010918. doi: 10.1371/journal.pcbi.1010918. eCollection 2023 Feb. PLoS Comput Biol. 2023. PMID: 36848395 Free PMC article.
-
Cutting Edge: Synapse Propensity of Human Memory CD8 T Cells Confers Competitive Advantage over Naive Counterparts.J Immunol. 2019 Aug 1;203(3):601-606. doi: 10.4049/jimmunol.1801687. Epub 2019 Jun 14. J Immunol. 2019. PMID: 31201237 Free PMC article.
References
-
- Bise R., Yin Z., Kanade T. Paper presented at: Biomedical Imaging: From Nano to Macro, 2011 IEEE International Symposium on (IEEE) 2011. Reliable cell tracking by global data association.
-
- Canny J. A computational approach to edge detection. IEEE Trans. Pattern Anal. Mach. Intell. 1986;679–698 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

