IFT27 links the BBSome to IFT for maintenance of the ciliary signaling compartment
- PMID: 25446516
- PMCID: PMC4254547
- DOI: 10.1016/j.devcel.2014.09.011
IFT27 links the BBSome to IFT for maintenance of the ciliary signaling compartment
Abstract
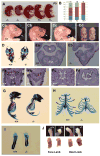
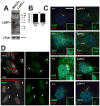
Vertebrate hedgehog signaling is coordinated by the differential localization of the receptors patched-1 and Smoothened in the primary cilium. Cilia assembly is mediated by intraflagellar transport (IFT), and cilia defects disrupt hedgehog signaling, causing many structural birth defects. We generated Ift25 and Ift27 knockout mice and show that they have structural birth defects indicative of hedgehog signaling dysfunction. Surprisingly, ciliary assembly is not affected, but abnormal hedgehog signaling is observed in conjunction with ciliary accumulation of patched-1 and Smoothened. Similarly, Smoothened accumulates in cilia on cells mutated for BBSome components or the BBS binding protein/regulator Lztfl1. Interestingly, the BBSome and Lztfl1 accumulate to high levels in Ift27 mutant cilia. Because Lztfl1 mutant cells accumulate BBSome but not IFT27, it is likely that Lztfl1 functions downstream of IFT27 to couple the BBSome to the IFT particle for coordinated removal of patched-1 and Smoothened from cilia during hedgehog signaling.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures





Comment in
-
Dissecting intraflagellar transport, one molecule at a time.Dev Cell. 2014 Nov 10;31(3):263-264. doi: 10.1016/j.devcel.2014.10.021. Epub 2014 Nov 10. Dev Cell. 2014. PMID: 25453827
Similar articles
-
Intraflagellar transport is deeply integrated in hedgehog signaling.Mol Biol Cell. 2018 May 15;29(10):1178-1189. doi: 10.1091/mbc.E17-10-0600. Epub 2018 Mar 22. Mol Biol Cell. 2018. PMID: 29540531 Free PMC article.
-
RABL4/IFT27 in a nucleotide-independent manner promotes phospholipase D ciliary retrieval via facilitating BBSome reassembly at the ciliary tip.J Cell Physiol. 2023 Mar;238(3):549-565. doi: 10.1002/jcp.30945. Epub 2023 Feb 28. J Cell Physiol. 2023. PMID: 36852649
-
Ubiquitin links smoothened to intraflagellar transport to regulate Hedgehog signaling.J Cell Biol. 2020 Jul 6;219(7):e201912104. doi: 10.1083/jcb.201912104. J Cell Biol. 2020. PMID: 32435793 Free PMC article.
-
Trafficking of ciliary membrane proteins by the intraflagellar transport/BBSome machinery.Essays Biochem. 2018 Dec 7;62(6):753-763. doi: 10.1042/EBC20180030. Print 2018 Dec 7. Essays Biochem. 2018. PMID: 30287585 Free PMC article. Review.
-
The roles of intraflagellar transport (IFT) protein 25 in mammalian signaling transduction and flagellogenesis.Asian J Androl. 2022 May-Jun;24(3):238-242. doi: 10.4103/aja202179. Asian J Androl. 2022. PMID: 34747727 Free PMC article. Review.
Cited by
-
Limited time window for retinal gene therapy in a preclinical model of ciliopathy.Hum Mol Genet. 2020 Aug 11;29(14):2337-2352. doi: 10.1093/hmg/ddaa124. Hum Mol Genet. 2020. PMID: 32568387 Free PMC article.
-
Microtubule Motors Drive Hedgehog Signaling in Primary Cilia.Trends Cell Biol. 2017 Feb;27(2):110-125. doi: 10.1016/j.tcb.2016.09.010. Epub 2016 Oct 17. Trends Cell Biol. 2017. PMID: 27765513 Free PMC article. Review.
-
Bardet-Biedl syndrome 3 regulates the development of cranial base midline structures.Bone. 2017 Aug;101:179-190. doi: 10.1016/j.bone.2016.02.017. Epub 2016 May 8. Bone. 2017. PMID: 27170093 Free PMC article.
-
The more we know, the more we have to discover: an exciting future for understanding cilia and ciliopathies.Cilia. 2015 Mar 31;4:5. doi: 10.1186/s13630-015-0014-0. eCollection 2015. Cilia. 2015. PMID: 25974046 Free PMC article.
-
Binding of IFT22 to the intraflagellar transport complex is essential for flagellum assembly.EMBO J. 2019 May 2;38(9):e101251. doi: 10.15252/embj.2018101251. Epub 2019 Apr 2. EMBO J. 2019. PMID: 30940671 Free PMC article.
References
-
- Aldahmesh MA, Li Y, Alhashem A, Anazi S, Alkuraya H, Hashem M, Awaji AA, Sogaty S, Alkharashi A, Alzahrani S, Al Hazzaa SA, Xiong Y, Kong S, Sun Z, Alkuraya FS. IFT27, encoding a small GTPase component of IFT particles, is mutated in a consanguineous family with Bardet-Biedl syndrome. Hum Mol Genet. 2014;23:3307–3315. - PMC - PubMed
-
- Badano JL, Mitsuma N, Beales PL, Katsanis N. The ciliopathies: an emerging class of human genetic disorders. Annu Rev Genomics Hum Genet. 2006;7:125–148. - PubMed
-
- Bai CB, Auerbach W, Lee JS, Stephen D, Joyner AL. Gli2, but not Gli1, is required for initial Shh signaling and ectopic activation of the Shh pathway. Development. 2002;129:4753–4761. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

