The MAPK ERK5, but not ERK1/2, inhibits the progression of monocytic phenotype to the functioning macrophage
- PMID: 25447310
- PMCID: PMC4364001
- DOI: 10.1016/j.yexcr.2014.10.003
The MAPK ERK5, but not ERK1/2, inhibits the progression of monocytic phenotype to the functioning macrophage
Abstract
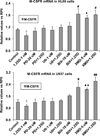
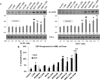
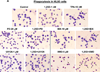
Intracellular signaling pathways present targets for pharmacological agents with potential for treatment of neoplastic diseases, with some disease remissions already recorded. However, cellular compensatory mechanisms usually negate the initial success. For instance, attempts to interrupt aberrant signaling downstream of the frequently mutated ras by inhibiting ERK1/2 has shown only limited usefulness for cancer therapy. Here, we examined how ERK5, that overlaps the functions of ERK1/2 in cell proliferation and survival, functions in a manner distinct from ERK1/2 in human AML cells induced to differentiate by 1,25D-dihydroxyvitamin D3 (1,25D). Using inhibitors of ERK1/2 and of MEK5/ERK5 at concentrations specific for each kinase in HL60 and U937 cells, we observed that selective inhibition of the kinase activity of ERK5, but not of ERK1/2, in the presence of 1,25D resulted in macrophage-like cell morphology and enhancement of phagocytic activity. Importantly, this was associated with increased expression of the macrophage colony stimulating factor receptor (M-CSFR), but was not seen when M-CSFR expression was knocked down. Interestingly, inhibition of ERK1/2 led to activation of ERK5 in these cells. Our results support the hypothesis that ERK5 negatively regulates the expression of M-CSFR, and thus has a restraining function on macrophage differentiation. The addition of pharmacological inhibitors of ERK5 may influence trials of differentiation therapy of AML.
Keywords: AML; ERK1/2; ERK5; M-CSFR; MAPK; Vitamin D.
Copyright © 2014 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors have no conflict of interest to disclose.
Figures
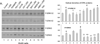
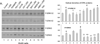







Similar articles
-
ERK1/2 and ERK5 have distinct roles in the regulation of brain-derived neurotrophic factor expression.J Neurosci Res. 2011 Oct;89(10):1542-50. doi: 10.1002/jnr.22683. Epub 2011 Jun 6. J Neurosci Res. 2011. PMID: 21647938
-
The ERK5 and ERK1/2 signaling pathways play opposing regulatory roles during chondrogenesis of adult human bone marrow-derived multipotent progenitor cells.J Cell Physiol. 2010 Jul;224(1):178-86. doi: 10.1002/jcp.22120. J Cell Physiol. 2010. PMID: 20232315
-
ERK 5/MAPK pathway has a major role in 1α,25-(OH)2 vitamin D3-induced terminal differentiation of myeloid leukemia cells.J Steroid Biochem Mol Biol. 2014 Oct;144 Pt A:223-7. doi: 10.1016/j.jsbmb.2013.10.002. Epub 2013 Oct 26. J Steroid Biochem Mol Biol. 2014. PMID: 24514755 Free PMC article. Review.
-
Retrograde neurotrophic signaling in rat retinal ganglion cells is transmitted via the ERK5 but not the ERK1/2 pathway.Invest Ophthalmol Vis Sci. 2014 Feb 3;55(2):658-65. doi: 10.1167/iovs.13-12985. Invest Ophthalmol Vis Sci. 2014. PMID: 24398098
-
MAPK signalling: ERK5 versus ERK1/2.EMBO Rep. 2006 Aug;7(8):782-6. doi: 10.1038/sj.embor.7400755. EMBO Rep. 2006. PMID: 16880823 Free PMC article. Review.
Cited by
-
Nuclear ERK5 inhibits progression of leukemic monocytes to macrophages by regulating the transcription factor PU.1 and heat shock protein HSP70.Leuk Lymphoma. 2017 Jun;58(6):1468-1480. doi: 10.1080/10428194.2016.1243675. Epub 2016 Oct 17. Leuk Lymphoma. 2017. PMID: 27748139 Free PMC article.
-
Prodifferentiation Activity of Novel Vitamin D₂ Analogs PRI-1916 and PRI-1917 and Their Combinations with a Plant Polyphenol in Acute Myeloid Leukemia Cells.Int J Mol Sci. 2016 Jul 5;17(7):1068. doi: 10.3390/ijms17071068. Int J Mol Sci. 2016. PMID: 27399677 Free PMC article.
-
The Potential of Vitamin D-Regulated Intracellular Signaling Pathways as Targets for Myeloid Leukemia Therapy.J Clin Med. 2015 Mar 25;4(4):504-34. doi: 10.3390/jcm4040504. J Clin Med. 2015. PMID: 26239344 Free PMC article. Review.
-
Molecular Mechanisms of Epithelial to Mesenchymal Transition Regulated by ERK5 Signaling.Biomolecules. 2021 Jan 29;11(2):183. doi: 10.3390/biom11020183. Biomolecules. 2021. PMID: 33572742 Free PMC article. Review.
-
Extracellular-Regulated Protein Kinase 5-Mediated Control of p21 Expression Promotes Macrophage Proliferation Associated with Tumor Growth and Metastasis.Cancer Res. 2020 Aug 15;80(16):3319-3330. doi: 10.1158/0008-5472.CAN-19-2416. Epub 2020 Jun 19. Cancer Res. 2020. PMID: 32561530 Free PMC article.
References
-
- Rademakers R, Baker M, Nicholson AM, Rutherford NJ, Finch N, Soto-Ortolaza A, Lash J, Wider C, Wojtas A, DeJesus-Hernandez M, Adamson J, Kouri N, Sundal C, Shuster EA, Aasly J, MacKenzie J, Roeber S, Kretzschmar HA, Boeve BF, Knopman DS, Petersen RC, Cairns NJ, Ghetti B, Spina S, Garbern J, Tselis AC, Uitti R, Das P, Van Gerpen JA, Meschia JF, Levy S, Broderick DF, Graff-Radford N, Ross OA, Miller BB, Swerdlow RH, Dickson DW, Wszolek ZK. Mutations in the colony stimulating factor 1 receptor (CSF1R) gene cause hereditary diffuse leukoencephalopathy with spheroids. Nature genetics. 2012;44:200–205. - PMC - PubMed
-
- Barreda DR, Hanington PC, Belosevic M. Regulation of myeloid development and function by colony stimulating factors. Developmental and comparative immunology. 2004;28:509–554. - PubMed
-
- Hamilton JA, Achuthan A. Colony stimulating factors and myeloid cell biology in health and disease. Trends in immunology. 2013;34:81–89. - PubMed
-
- Broughton SE, Hercus TR, Lopez AF, Parker MW. Cytokine receptor activation at the cell surface. Current opinion in structural biology. 2012;22:350–359. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

