Nucleocytoplasmic shuttling of valosin-containing protein (VCP/p97) regulated by its N domain and C-terminal region
- PMID: 25447673
- PMCID: PMC4254625
- DOI: 10.1016/j.bbamcr.2014.10.019
Nucleocytoplasmic shuttling of valosin-containing protein (VCP/p97) regulated by its N domain and C-terminal region
Abstract
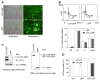
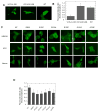
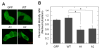
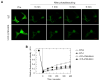
Valosin-containing protein (VCP or p97), a member of the AAA family (ATPases associated with diverse cellular activities), plays a key role in many important cellular activities. A genetic deficiency of VCP can cause inclusion body myopathy associated with Paget's disease of bone and frontotemporal dementia (IBMPFD). Previous studies showed that the VCP N domain is essential for the regulation of nuclear entry of VCP. Here we report that IBMPFD mutations, which are mainly located in the N domain, suppress the nuclear entry of VCP. Moreover, the peptide sequence G780AGPSQ in the C-terminal region regulates the retention of VCP in the nucleus. A mutant lacking this sequence can increase the nuclear distribution of IBMPFD VCP, suggesting that this sequence is a potential molecular target for correcting the deficient nucleocytoplasmic shuttling of IBMPFD VCP proteins.
Keywords: Inclusion body myopathy associated with Paget's disease of bone and frontotemporal dementia (IBMPFD); Nuclear export signal; Nucleocytoplasmic shuttling; Valosin containing protein.
Copyright © 2014 Elsevier B.V. All rights reserved.
Figures



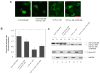

References
-
- Wang Q, Song C, Li CC. Molecular perspectives on p97-VCP: progress in understanding its structure and diverse biological functions. J Struct Biol. 2004;146:44–57. - PubMed
-
- Meyer H, Bug M, Bremer S. Emerging functions of the VCP/p97 AAA-ATPase in the ubiquitin system. Nat Cell Biol. 2012;14:117–123. - PubMed
-
- Yamanaka K, Sasagawa Y, Ogura T. Recent advances in p97/VCP/Cdc48 cellular functions. Biochim Biophys Acta. 2012;1823:130–137. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

