Rapid prototyping of microbial cell factories via genome-scale engineering
- PMID: 25450192
- PMCID: PMC4439387
- DOI: 10.1016/j.biotechadv.2014.11.007
Rapid prototyping of microbial cell factories via genome-scale engineering
Abstract
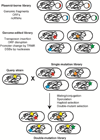
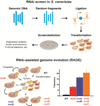
Advances in reading, writing and editing genetic materials have greatly expanded our ability to reprogram biological systems at the resolution of a single nucleotide and on the scale of a whole genome. Such capacity has greatly accelerated the cycles of design, build and test to engineer microbes for efficient synthesis of fuels, chemicals and drugs. In this review, we summarize the emerging technologies that have been applied, or are potentially useful for genome-scale engineering in microbial systems. We will focus on the development of high-throughput methodologies, which may accelerate the prototyping of microbial cell factories.
Keywords: Genome synthesis; Genome-scale engineering; High-throughput technology; Homologous recombination; Microbial cell factory; Transcriptome engineering.
Copyright © 2014 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures

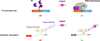
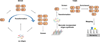
References
-
- Albert H, Dale EC, Lee E, Ow DW. Site-specific integration of DNA into wild-type and mutant lox sites placed in the plant genome. Plant J. 1995;7:649–659. - PubMed
-
- Alexeyev MF, Shokolenko IN. Mini-Tn10 transposon derivatives for insertion mutagenesis and gene delivery into the chromosome of gram-negative bacteria. Gene. 1995;160:59–62. - PubMed
-
- Alper H, Moxley J, Nevoigt E, Fink GR, Stephanopoulos G. Engineering yeast transcription machinery for improved ethanol tolerance and production. Science. 2006;314:1565–1568. - PubMed
-
- Alper H, Stephanopoulos G. Global transcription machinery engineering: a new approach for improving cellular phenotype. Metab Eng. 2007;9:258–267. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

