Conserved higher-order chromatin regulates NMDA receptor gene expression and cognition
- PMID: 25467983
- PMCID: PMC4258154
- DOI: 10.1016/j.neuron.2014.10.032
Conserved higher-order chromatin regulates NMDA receptor gene expression and cognition
Abstract
Three-dimensional chromosomal conformations regulate transcription by moving enhancers and regulatory elements into spatial proximity with target genes. Here we describe activity-regulated long-range loopings bypassing up to 0.5 Mb of linear genome to modulate NMDA glutamate receptor GRIN2B expression in human and mouse prefrontal cortex. Distal intronic and 3' intergenic loop formations competed with repressor elements to access promoter-proximal sequences, and facilitated expression via a "cargo" of AP-1 and NRF-1 transcription factors and TALE-based transcriptional activators. Neuronal deletion or overexpression of Kmt2a/Mll1 H3K4- and Kmt1e/Setdb1 H3K9-methyltransferase was associated with higher-order chromatin changes at distal regulatory Grin2b sequences and impairments in working memory. Genetic polymorphisms and isogenic deletions of loop-bound sequences conferred liability for cognitive performance and decreased GRIN2B expression. Dynamic regulation of chromosomal conformations emerges as a novel layer for transcriptional mechanisms impacting neuronal signaling and cognition.
Copyright © 2014 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors thank the participants and the personnel of the Military Training Camp of Candidate, Supply Army officers (SEAP) in Heraklion, Crete for their help with the LOGOS study. The authors report no conflicts.
Figures




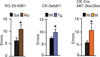
References
-
- Banerji J, Rusconi S, Schaffner W. Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell. 1981;27:299–308. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

