A novel biomarker Linc00974 interacting with KRT19 promotes proliferation and metastasis in hepatocellular carcinoma
- PMID: 25476897
- PMCID: PMC4649834
- DOI: 10.1038/cddis.2014.518
A novel biomarker Linc00974 interacting with KRT19 promotes proliferation and metastasis in hepatocellular carcinoma
Abstract
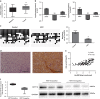
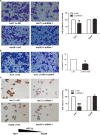
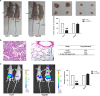
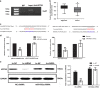
Location-associated long noncoding RNA (lncRNA) was reported to interact with target protein via a cis-regulatory process especially for the Flank10kb class lncRNA. Based on this theory, we aimed to explore the regulatory mechanisms of Linc00974 and KRT19 (an lncRNA beyond the Flank10kb class with protein) when we first confirmed the aberrant expression in hepatocellular carcinoma in a previous study. Knockdown of Linc00974 resulted in an inhibition of cell proliferation and invasion with an activation of apoptosis and cell cycle arrest in vitro, which was also validated by a subcutaneous and tail vein/intraperitoneal injection xenotransplantation model in vivo. We further investigated the interaction pattern of Linc00974 and KRT19. MiR-642 was identified, by acting as the competing endogenous RNA in regulating Linc00974 and KRT19. Linc00974 was increased owing to an abnormal hypomethylation promoter, which induced the upregulation of KRT19 via ceRNA interaction, resulting in the activation of the Notch and TGF-β pathways as detected by cDNA microarray. We also discovered Linc00974F-1 stably expressed in the plasma. By the combined analysis of Linc00974F-1 with CYFRA21-1, we found that these joint indicators predicted growth and metastasis of tumor in HCC patients. In conclusion, the combination of Linc00974 and KRT19 may be novel indices for clinical diagnosis of tumor growth and metastasis in HCC, while Linc00974 may become a potential therapeutic target for the prevention of HCC progression.
Figures

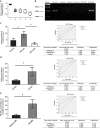
References
-
- 1Shen Q, Fan J, Yang XR, Tan Y, Zhao W, Xu Y et al. Serum DKK1 as a protein biomarker for the diagnosis of hepatocellular carcinoma: a large-scale, multicentre study. Lancet Oncol 2012; 13: 817–826. - PubMed
-
- 5Xu D, Yang F, Yuan JH, Zhang L, Bi HS, Zhou CC et al. Long noncoding RNAs associated with liver regeneration 1 accelerates hepatocyte proliferation during liver regeneration by activating Wnt/beta-catenin signaling. Hepatology 2013; 58: 739–751. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

