Systems analysis of high-throughput data
- PMID: 25480641
- PMCID: PMC4426208
- DOI: 10.1007/978-1-4939-2095-2_8
Systems analysis of high-throughput data
Abstract
Modern high-throughput assays yield detailed characterizations of the genomic, transcriptomic, and proteomic states of biological samples, enabling us to probe the molecular mechanisms that regulate hematopoiesis or give rise to hematological disorders. At the same time, the high dimensionality of the data and the complex nature of biological interaction networks present significant analytical challenges in identifying causal variations and modeling the underlying systems biology. In addition to identifying significantly disregulated genes and proteins, integrative analysis approaches that allow the investigation of these single genes within a functional context are required. This chapter presents a survey of current computational approaches for the statistical analysis of high-dimensional data and the development of systems-level models of cellular signaling and regulation. Specifically, we focus on multi-gene analysis methods and the integration of expression data with domain knowledge (such as biological pathways) and other gene-wise information (e.g., sequence or methylation data) to identify novel functional modules in the complex cellular interaction network.
Figures
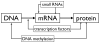
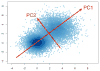
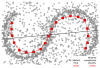



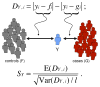




References
-
- van den Akker-van Marle ME, Gurwitz D, Detmar SB, Enzing CM, Hopkins MM, de Mesa EG, Ibarreta D. Cost-effectiveness of pharmacogenomics in clinical practice: a case study of thiopurine methyltransferase genotyping in acute lymphoblastic leukemia in Europe. Pharmacogenomics. 2006;7(5):783–92. - PubMed
-
- Karajannis M, Vincent L, Direnzo R, Shmelkov S, Zhang F, Feldman E, Bohlen P, Zhu Z, Sun H, Kussie P, Rafii S. Activation of fgfr1beta signaling pathway promotes survival, migration and resistance to chemotherapy in acute myeloid leukemia cells. Leukemia. 2006 - PubMed
-
- Savageau MA, Rosen R. Biochemical systems analysis: a study of function and design in molecular biology. Vol. 725. Addison-Wesley; Reading, MA: 1976.
-
- Von Bertalanffy L. In: Modern theories of development: An introduction to theoretical biology. Woodger JH, translator. Oxford University Press; 1933. originally published 1928.
-
- Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4(2):249–264. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

