Isothermal chemical denaturation to determine binding affinity of small molecules to G-protein coupled receptors
- PMID: 25481736
- PMCID: PMC4355058
- DOI: 10.1016/j.ab.2014.11.019
Isothermal chemical denaturation to determine binding affinity of small molecules to G-protein coupled receptors
Abstract
The determination of accurate binding affinities is critical in drug discovery and development. Several techniques are available for characterizing the binding of small molecules to soluble proteins. The situation is different for integral membrane proteins. Isothermal chemical denaturation has been shown to be a valuable biophysical method to determine, in a direct and label-free fashion, the binding of ligands to soluble proteins. In this study, the application of isothermal chemical denaturation was applied to an integral membrane protein, the A2a G-protein coupled receptor. Binding affinities for a set of 19 small molecule agonists/antagonists of the A2a receptor were determined and found to be in agreement with data from surface plasmon resonance and radioligand binding assays previously reported in the literature. Therefore, isothermal chemical denaturation expands the available toolkit of biophysical techniques to characterize and study ligand binding to integral membrane proteins, specifically G-protein coupled receptors in vitro.
Keywords: Binding affinity determination; Binding thermodynamics; Chemical denaturation; GPCR.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures
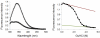



Similar articles
-
Real-time monitoring of binding events on a thermostabilized human A2A receptor embedded in a lipid bilayer by surface plasmon resonance.Biochim Biophys Acta. 2015 May;1848(5):1224-33. doi: 10.1016/j.bbamem.2015.02.014. Epub 2015 Feb 25. Biochim Biophys Acta. 2015. PMID: 25725488
-
Bridging molecular docking to membrane molecular dynamics to investigate GPCR-ligand recognition: the human A₂A adenosine receptor as a key study.J Chem Inf Model. 2014 Jan 27;54(1):169-83. doi: 10.1021/ci400532b. Epub 2014 Jan 8. J Chem Inf Model. 2014. PMID: 24359090
-
Integrating Pharmacophore into Membrane Molecular Dynamics Simulations to Improve Homology Modeling of G Protein-coupled Receptors with Ligand Selectivity: A2A Adenosine Receptor as an Example.Chem Biol Drug Des. 2015 Dec;86(6):1438-50. doi: 10.1111/cbdd.12607. Epub 2015 Jul 14. Chem Biol Drug Des. 2015. PMID: 26072970
-
Adenosine A2A receptor as a drug discovery target.J Med Chem. 2014 May 8;57(9):3623-50. doi: 10.1021/jm4011669. Epub 2013 Nov 15. J Med Chem. 2014. PMID: 24164628 Review.
-
Novel approaches for targeting the adenosine A2A receptor.Expert Opin Drug Discov. 2015 Jan;10(1):63-80. doi: 10.1517/17460441.2015.971006. Epub 2014 Oct 14. Expert Opin Drug Discov. 2015. PMID: 25311639 Review.
Cited by
-
Stabilization of G protein-coupled receptors by point mutations.Front Pharmacol. 2015 Apr 20;6:82. doi: 10.3389/fphar.2015.00082. eCollection 2015. Front Pharmacol. 2015. PMID: 25941489 Free PMC article. Review.
-
Fluorescence-Based Protein Stability Monitoring-A Review.Int J Mol Sci. 2024 Feb 1;25(3):1764. doi: 10.3390/ijms25031764. Int J Mol Sci. 2024. PMID: 38339045 Free PMC article. Review.
References
-
- Navratilova I, Dioszegi M, Myszka DG. Analyzing ligand and small molecule binding activity of solubilized GPCRs using biosensor technology. Anal Biochem. 2006;355:132–139. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

