Electrostatic interactions between the Bni1p Formin FH2 domain and actin influence actin filament nucleation
- PMID: 25482541
- PMCID: PMC4286494
- DOI: 10.1016/j.str.2014.10.014
Electrostatic interactions between the Bni1p Formin FH2 domain and actin influence actin filament nucleation
Abstract
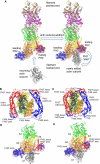
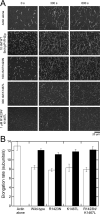
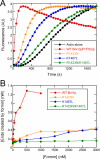
Formins catalyze nucleation and growth of actin filaments. Here, we study the structure and interactions of actin with the FH2 domain of budding yeast formin Bni1p. We built an all-atom model of the formin dimer on an Oda actin filament 7-mer and studied structural relaxation and interprotein interactions by molecular dynamics simulations. These simulations produced a refined model for the FH2 dimer associated with the barbed end of the filament and showed electrostatic interactions between the formin knob and actin target-binding cleft. Mutations of two formin residues contributing to these interactions (R1423N, K1467L, or both) reduced the interaction energies between the proteins, and in coarse-grained simulations, the formin lost more interprotein contacts with an actin dimer than with an actin 7-mer. Biochemical experiments confirmed a strong influence of these mutations on Bni1p-mediated actin filament nucleation, but not elongation, suggesting that different interactions contribute to these two functions of formins.
Copyright © 2015 Elsevier Ltd. All rights reserved.
Figures





References
-
- Brown WM, Kohlmeyer A, Plimpton SJ, Tharrington AN. Implementing molecular dynamics on hybrid high performance computers - Particle-particle particle-mesh. Comput Phys Commun. 2012;183:449–459.
-
- Brown WM, Wang P, Plimpton SJ, Tharrington AN. Implementing molecular dynamics on hybrid high performance computers - short range forces. Comput Phys Commun. 2011;182:898–911.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

