The role of proteases in regulating Eph/ephrin signaling
- PMID: 25482632
- PMCID: PMC4594553
- DOI: 10.4161/19336918.2014.970026
The role of proteases in regulating Eph/ephrin signaling
Abstract
Proteases regulate a myriad of cell functions, both in normal and disease states. In addition to protein turnover, they regulate a range of signaling processes, including those mediated by Eph receptors and their ephrin ligands. A variety of proteases is reported to directly cleave Ephs and/or ephrins under different conditions, to promote receptor and/or ligand shedding, and regulate receptor/ligand internalisation and signaling. They also cleave other adhesion proteins in response to Eph-ephrin interactions, to indirectly facilitate Eph-mediated functions. Proteases thus contribute to Eph/ephrin mediated changes in cell-cell and cell-matrix interactions, in cell morphology and in cell migration and invasion, in a manner which appears to be tightly regulated by, and co-ordinated with, Eph signaling. This review summarizes the current literature describing the function and regulation of protease activities during Eph/ephrin-mediated cell signaling.
Keywords: AD, Alzheimer's disease; ADAM, a disintegrin and metalloprotease; APP, Amyloid precursor protein; Arf1, ADP rybosylation factor 1; BACE, β-site APP cleaving enzyme; CAM, cell adhesion molecules; CTF, cytoplasmic fragment; CNC, cranial neural crest; ECM,; ADAM; Eph receptor tyrosine kinase; MMP; ephrin signaling; protease; γ-secretase.
Figures
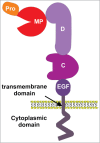
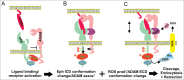
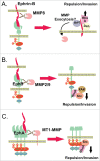

References
-
- Serim S, Haedke U, Verhelst SHL. Activity-based probes for the study of proteases: recent advances and developments. ChemMedChem 2012; 7:1146-59; PMID:22431376; http://dx.doi.org/ 10.1002/cmdc.201200057 - DOI - PubMed
-
- Drag M, Salvesen GS. Emerging principles in protease-based drug discovery. Nat Rev Drug Discov 2010; 9:690-701; PMID:20811381; http://dx.doi.org/ 10.1038/nrd3053 - DOI - PMC - PubMed
-
- Turk B. Targeting proteases: successes, failures and future prospects. Nat Rev Drug Dis 2006; 5:785-99; PMID:16955069; http://dx.doi.org/ 10.1038/nrd2092 - DOI - PubMed
-
- Lopez-Otin C, Bond JS. Proteases: multifunctional enzymes in life and disease. J Biol Chem 2008; 283:30433-7; PMID:18650443; http://dx.doi.org/ 10.1074/jbc.R800035200 - DOI - PMC - PubMed
-
- Overall CM, Blobel CP. In search of partners: linking extracellular proteases to substrates. Nat Rev Mol Cell Biol 2007; 8:245-57; PMID:17299501; http://dx.doi.org/ 10.1038/nrm2120 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
