Proteins involved in the degradation of cytoplasmic mRNA in the major eukaryotic model systems
- PMID: 25483043
- PMCID: PMC4615280
- DOI: 10.4161/rna.34406
Proteins involved in the degradation of cytoplasmic mRNA in the major eukaryotic model systems
Abstract
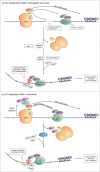
The process of mRNA decay and surveillance is considered to be one of the main posttranscriptional gene expression regulation platforms in eukaryotes. The degradation of stable, protein-coding transcripts is normally initiated by removal of the poly(A) tail followed by 5'-cap hydrolysis and degradation of the remaining mRNA body by Xrn1. Alternatively, the exosome complex degrades mRNA in the 3'>5'direction. The newly discovered uridinylation-dependent pathway, which is present in many different organisms, also seems to play a role in bulk mRNA degradation. Simultaneously, to avoid the synthesis of incorrect proteins, special cellular machinery is responsible for the removal of faulty transcripts via nonsense-mediated, no-go, non-stop or non-functional 18S rRNA decay. This review is focused on the major eukaryotic cytoplasmic mRNA degradation pathways showing many similarities and pointing out main differences between the main model-species: yeast, Drosophila, plants and mammals.
Keywords: Arabidopsis thaliana; Drosophila melanogaster; Saccharomyces cerevisiae; Schizosaccharomyces pombe; human; mRNA decay; mRNA surveillance.
Figures



Similar articles
-
Cytoplasmic RNA decay pathways - Enzymes and mechanisms.Biochim Biophys Acta. 2016 Dec;1863(12):3125-3147. doi: 10.1016/j.bbamcr.2016.09.023. Epub 2016 Oct 3. Biochim Biophys Acta. 2016. PMID: 27713097 Review.
-
Messenger RNA turnover in eukaryotes: pathways and enzymes.Crit Rev Biochem Mol Biol. 2004 Jul-Aug;39(4):197-216. doi: 10.1080/10409230490513991. Crit Rev Biochem Mol Biol. 2004. PMID: 15596551 Review.
-
[Destroy this (RNA) message after reading it!].Med Sci (Paris). 2007 Oct;23(10):850-6. doi: 10.1051/medsci/20072310850. Med Sci (Paris). 2007. PMID: 17937894 Review. French.
-
mRNA decay: x (XRN1) marks the spot.Mol Cell. 2003 May;11(5):1126-8. doi: 10.1016/s1097-2765(03)00198-9. Mol Cell. 2003. PMID: 12769838 Review.
-
The eukaryotic transcriptional machinery regulates mRNA translation and decay in the cytoplasm.Biochim Biophys Acta. 2013 Jan;1829(1):169-73. doi: 10.1016/j.bbagrm.2012.08.004. Epub 2012 Sep 6. Biochim Biophys Acta. 2013. PMID: 22982191 Review.
Cited by
-
Long-term durability of immune responses to the BNT162b2 and mRNA-1273 vaccines based on dosage, age and sex.Sci Rep. 2022 Dec 8;12(1):21232. doi: 10.1038/s41598-022-25134-0. Sci Rep. 2022. PMID: 36481777 Free PMC article.
-
Immunoglobulin expression and the humoral immune response is regulated by the non-canonical poly(A) polymerase TENT5C.Nat Commun. 2020 Apr 27;11(1):2032. doi: 10.1038/s41467-020-15835-3. Nat Commun. 2020. PMID: 32341344 Free PMC article.
-
The rates of the major steps in the molecular mechanism of RNase H1-dependent antisense oligonucleotide induced degradation of RNA.Nucleic Acids Res. 2015 Oct 15;43(18):8955-63. doi: 10.1093/nar/gkv920. Epub 2015 Sep 17. Nucleic Acids Res. 2015. PMID: 26384424 Free PMC article.
-
Nonsense-mediated mRNA decay: an intricate machinery that shapes transcriptomes.Nat Rev Mol Cell Biol. 2015 Nov;16(11):665-77. doi: 10.1038/nrm4063. Epub 2015 Sep 23. Nat Rev Mol Cell Biol. 2015. PMID: 26397022 Review.
-
Perlman syndrome nuclease DIS3L2 controls cytoplasmic non-coding RNAs and provides surveillance pathway for maturing snRNAs.Nucleic Acids Res. 2016 Dec 1;44(21):10437-10453. doi: 10.1093/nar/gkw649. Epub 2016 Jul 18. Nucleic Acids Res. 2016. PMID: 27431325 Free PMC article.
References
-
- Garneau NL, Wilusz J, Wilusz CJ. The highways and byways of mRNA decay. Nat Rev Mol Cell Biol 2007; 8:113-26; PMID:17245413; http://dx.doi.org/10.1038/nrm2104 - DOI - PubMed
-
- Chen C-YA, Shyu A-B. Protein segregase meddles in remodeling of mRNA-protein complexes. Genes Dev 2013; 27:980-4; PMID:23651853; http://dx.doi.org/10.1101/gad.219469.113 - DOI - PMC - PubMed
-
- Barreau C, Paillard L, Osborne HB. AU-rich elements and associated factors: are there unifying principles? Nucleic Acids Res 2005; 33:7138-50; PMID:16391004; http://dx.doi.org/10.1093/nar/gki1012 - DOI - PMC - PubMed
-
- Chen C-YA, Shyu A-B. Rapid deadenylation triggered by a nonsense codon precedes decay of the RNA body in a mammalian cytoplasmic nonsense-mediated decay pathway. Mol Cell Biol 2003; 23:4805-13; PMID:12832468; http://dx.doi.org/10.1128/MCB.23.14.4805-4813.2003 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
