Specificity rendering 'hot-spots' for aurora kinase inhibitor design: the role of non-covalent interactions and conformational transitions
- PMID: 25485544
- PMCID: PMC4259475
- DOI: 10.1371/journal.pone.0113773
Specificity rendering 'hot-spots' for aurora kinase inhibitor design: the role of non-covalent interactions and conformational transitions
Abstract
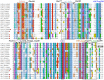
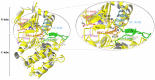
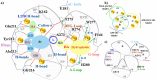
The present study examines the conformational transitions occurring among the major structural motifs of Aurora kinase (AK) concomitant with the DFG-flip and deciphers the role of non-covalent interactions in rendering specificity. Multiple sequence alignment, docking and structural analysis of a repertoire of 56 crystal structures of AK from Protein Data Bank (PDB) has been carried out. The crystal structures were systematically categorized based on the conformational disposition of the DFG-loop [in (DI) 42, out (DO) 5 and out-up (DOU) 9], G-loop [extended (GE) 53 and folded (GF) 3] and αC-helix [in (CI) 42 and out (CO) 14]. The overlapping subsets on categorization show the inter-dependency among structural motifs. Therefore, the four distinct possibilities a) 2W1C (DI, CI, GE) b) 3E5A (DI, CI, GF) c) 3DJ6 (DI, CO, GF) d) 3UNZ (DOU, CO, GF) along with their co-crystals and apo-forms were subjected to molecular dynamics simulations of 40 ns each to evaluate the variations of individual residues and their impact on forming interactions. The non-covalent interactions formed by the 157 AK co-crystals with different regions of the binding site were initially studied with the docked complexes and structure interaction fingerprints. The frequency of the most prominent interactions was gauged in the AK inhibitors from PDB and the four representative conformations during 40 ns. Based on this study, seven major non-covalent interactions and their complementary sites in AK capable of rendering specificity have been prioritized for the design of different classes of inhibitors.
Conflict of interest statement
Figures



Similar articles
-
Modeling conformational flexibility of kinases in inactive states.Proteins. 2019 Nov;87(11):943-951. doi: 10.1002/prot.25756. Epub 2019 Jun 17. Proteins. 2019. PMID: 31168936 Free PMC article.
-
Quantitative conformational profiling of kinase inhibitors reveals origins of selectivity for Aurora kinase activation states.Proc Natl Acad Sci U S A. 2018 Dec 18;115(51):E11894-E11903. doi: 10.1073/pnas.1811158115. Epub 2018 Dec 5. Proc Natl Acad Sci U S A. 2018. PMID: 30518564 Free PMC article.
-
Structural elucidation of the DFG-Asp in and DFG-Asp out states of TAM kinases and insight into the selectivity of their inhibitors.Molecules. 2014 Oct 10;19(10):16223-39. doi: 10.3390/molecules191016223. Molecules. 2014. PMID: 25310149 Free PMC article.
-
Structural Biology Insight for the Design of Sub-type Selective Aurora Kinase Inhibitors.Curr Cancer Drug Targets. 2015;15(5):375-93. doi: 10.2174/1568009615666150421110401. Curr Cancer Drug Targets. 2015. PMID: 25895501 Review.
-
The ABC of protein kinase conformations.Biochim Biophys Acta. 2015 Oct;1854(10 Pt B):1555-66. doi: 10.1016/j.bbapap.2015.03.009. Epub 2015 Apr 1. Biochim Biophys Acta. 2015. PMID: 25839999 Review.
Cited by
-
Lead identification for the K-Ras protein: virtual screening and combinatorial fragment-based approaches.Onco Targets Ther. 2016 May 2;9:2575-84. doi: 10.2147/OTT.S99671. eCollection 2016. Onco Targets Ther. 2016. PMID: 27217775 Free PMC article.
-
Theoretical investigation into the cooperativity effect between the intermolecular π∙π and H-bonding interactions in the curcumin∙cytosine∙H2O system.J Mol Model. 2018 Sep 28;24(10):298. doi: 10.1007/s00894-018-3836-z. J Mol Model. 2018. PMID: 30267159
-
Dynamics of human protein kinase Aurora A linked to drug selectivity.Elife. 2018 Jun 14;7:e36656. doi: 10.7554/eLife.36656. Elife. 2018. PMID: 29901437 Free PMC article.
-
Protein-protein interaction of RdRp with its co-factor NSP8 and NSP7 to decipher the interface hotspot residues for drug targeting: A comparison between SARS-CoV-2 and SARS-CoV.J Mol Struct. 2022 Jun 5;1257:132602. doi: 10.1016/j.molstruc.2022.132602. Epub 2022 Feb 8. J Mol Struct. 2022. PMID: 35153334 Free PMC article.
-
Towards systematic exploration of chemical space: building the fragment library module in molecular property diagnostic suite.Mol Divers. 2023 Jun;27(3):1459-1468. doi: 10.1007/s11030-022-10506-5. Epub 2022 Aug 4. Mol Divers. 2023. PMID: 35925528 Free PMC article.
References
-
- Yan A, Wang L, Xu S, Xu J (2011) Aurora-A kinase inhibitor scaffolds and binding modes. Drug Discov Today 16:260–269. - PubMed
-
- Nigg EA (2001) Mitotic kinases as regulators of cell division and its checkpoints. Nat Rev Mol Cell Biol 2:21–32. - PubMed
-
- Mountzios G, Terpos E, Dimopoulos MA (2008) Aurora kinases as targets for cancer therapy. Cancer Treat Rev 34:175–182. - PubMed
-
- Libertini S, Abagnale A, Passaro C, Botta G, Portella G (2010) Aurora A and B kinases-targets of novel anticancer drugs. Recent Pat Anticancer Drug Discov 5:219–241. - PubMed
-
- Harrington EA, Bebbington D, Moore J, Rasmussen RK, Ajose-Adeogun AO, et al. (2004) VX-680, a potent and selective small-molecule inhibitor of the Aurora kinases, suppresses tumor growth in vivo. Nat Med10:262–267. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

