Structural templates for comparative protein docking
- PMID: 25488330
- PMCID: PMC4461561
- DOI: 10.1002/prot.24736
Structural templates for comparative protein docking
Abstract
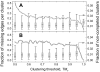
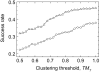
Structural characterization of protein-protein interactions is important for understanding life processes. Because of the inherent limitations of experimental techniques, such characterization requires computational approaches. Along with the traditional protein-protein docking (free search for a match between two proteins), comparative (template-based) modeling of protein-protein complexes has been gaining popularity. Its development puts an emphasis on full and partial structural similarity between the target protein monomers and the protein-protein complexes previously determined by experimental techniques (templates). The template-based docking relies on the quality and diversity of the template set. We present a carefully curated, nonredundant library of templates containing 4950 full structures of binary complexes and 5936 protein-protein interfaces extracted from the full structures at 12 Å distance cut-off. Redundancy in the libraries was removed by clustering the PDB structures based on structural similarity. The value of the clustering threshold was determined from the analysis of the clusters and the docking performance on a benchmark set. High structural quality of the interfaces in the template and validation sets was achieved by automated procedures and manual curation. The library is included in the Dockground resource for molecular recognition studies at http://dockground.bioinformatics.ku.edu.
Keywords: benchmark sets; protein interactions; protein modeling; protein recognition; structure prediction.
© 2014 Wiley Periodicals, Inc.
Figures





References
-
- Aloy P, Pichaud M, Russell RB. Protein complexes: Structure prediction challenges for the 21st century. Curr Opin Struct Biol. 2005;15:15–22. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

