Overexpression of tomato SlNAC1 transcription factor alters fruit pigmentation and softening
- PMID: 25491370
- PMCID: PMC4272553
- DOI: 10.1186/s12870-014-0351-y
Overexpression of tomato SlNAC1 transcription factor alters fruit pigmentation and softening
Abstract
Background: Fruit maturation and ripening are genetically regulated processes that involve a complex interplay of plant hormones, growth regulators and multiple biological and environmental factors. Tomato (Solanum lycopersicum) has been used as a model of biological and genetic studies on the regulation of specific ripening pathways, including ethylene, carotenoid and cell wall metabolism. This model has also been used to investigate the functions of upstream signalling and transcriptional regulators. Thus far, many ripening-associated transcription factors that influence fruit development and ripening have been reported. NAC transcription factors are plant specific and play important roles in many stages of plant growth and development, such as lateral root formation, secondary cell wall synthesis, and embryo, floral organ, vegetative organ and fruit development.
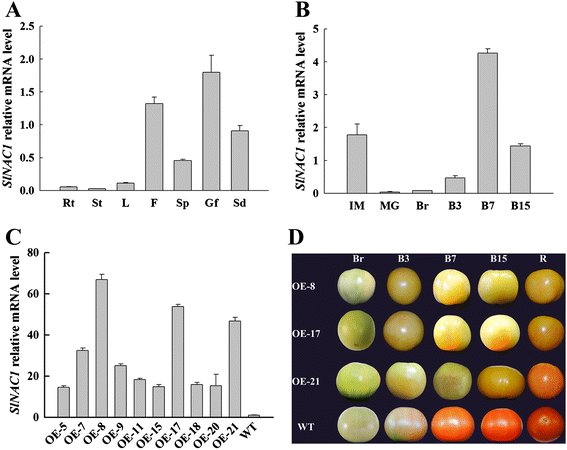
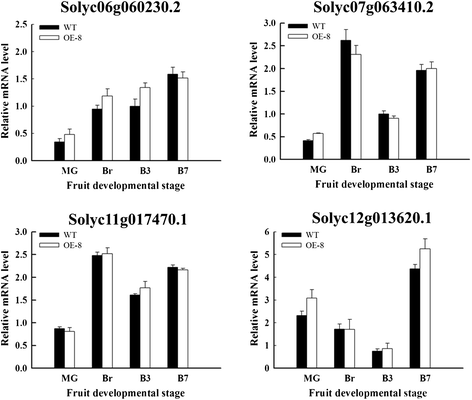
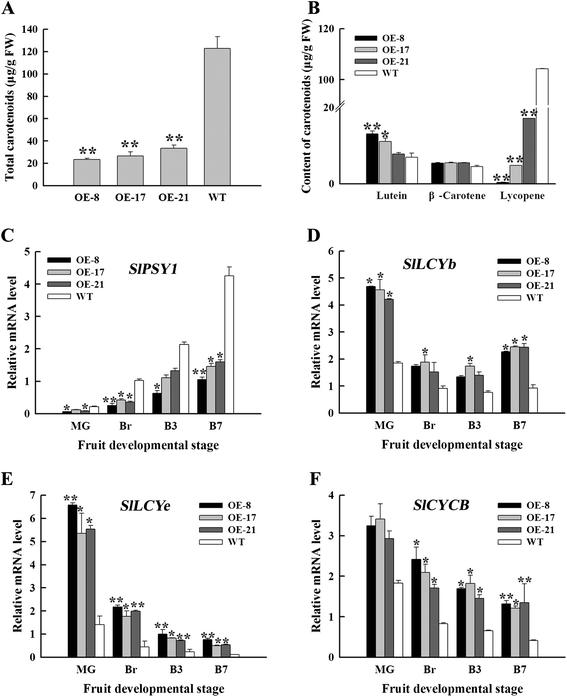
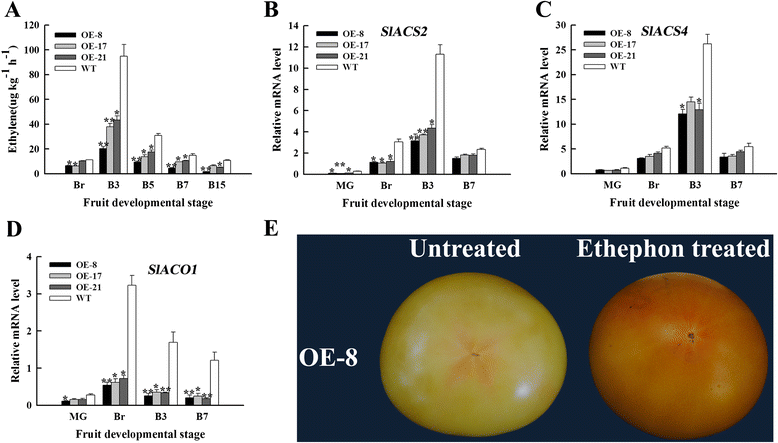
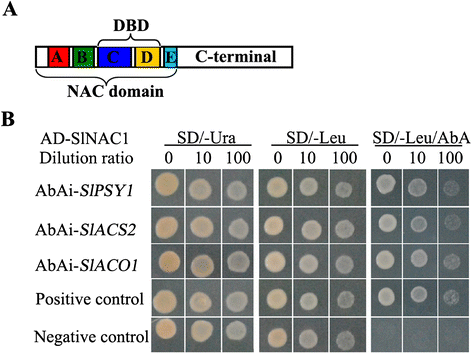
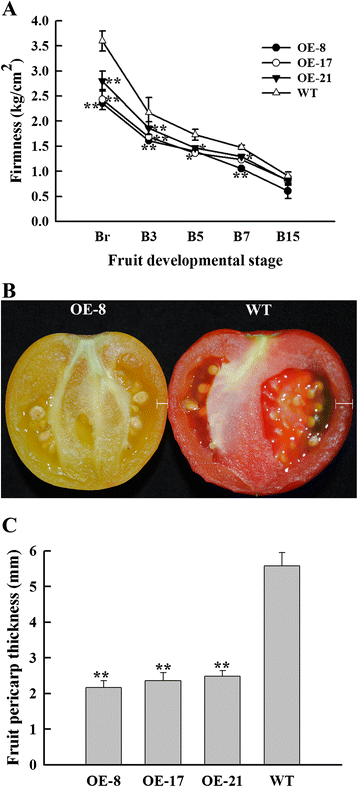
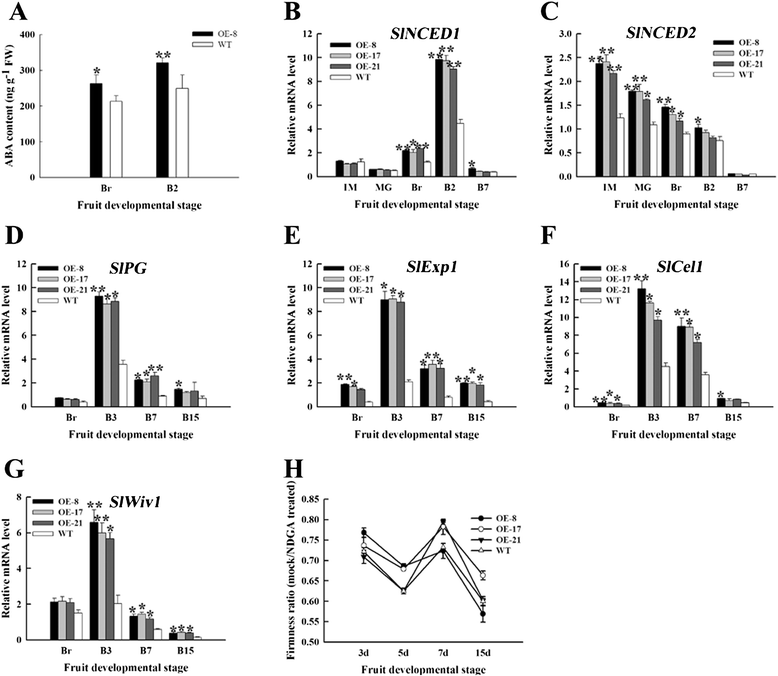
Results: Tissue-specific analysis by quantitative real-time PCR showed that SlNAC1 was highly accumulated in immature green fruits; the expression of SlNAC1 increased with fruit ripening till to the highest level at 7 d after the breaker stage. The overexpression of SlNAC1 resulted in reduced carotenoids by altering carotenoid pathway flux and decreasing ethylene synthesis mediated mainly by the reduced expression of ethylene biosynthetic genes of system-2, thus led to yellow or orange mature fruits. The results of yeast one-hybrid experiment demonstrated that SlNAC1 can interact with the regulatory regions of genes related lycopene and ethylene synthesis. These results also indicated that SlNAC1 inhibited fruit ripening by affecting ethylene synthesis and carotenoid accumulation in SlNAC1 overexpression lines. In addition, the overexpression of SlNAC1 reduced the firmness of the fruits and the thickness of the pericarp and produced more abscisic acid, resulting in the early softening of fruits. Hence, in SlNAC1 overexpression lines, both ethylene-dependent and abscisic acid-dependent pathways are regulated by SlNAC1 in fruit ripening regulatory network.
Conclusions: SlNAC1 had a broad influence on tomato fruit ripening and regulated SlNAC1 overexpression tomato fruit ripening through both ethylene-dependent and abscisic acid-dependent pathways. Thus, this study provided new insights into the current model of tomato fruit ripening regulatory network.
Figures
Similar articles
-
Suppression of tomato SlNAC1 transcription factor delays fruit ripening.J Plant Physiol. 2016 Apr 1;193:88-96. doi: 10.1016/j.jplph.2016.01.014. Epub 2016 Feb 26. J Plant Physiol. 2016. PMID: 26962710
-
SlERF109-like and SlNAC1 Coordinately Regulated Tomato Ripening by Inhibiting ACO1 Transcription.Int J Mol Sci. 2024 Feb 3;25(3):1873. doi: 10.3390/ijms25031873. Int J Mol Sci. 2024. PMID: 38339150 Free PMC article.
-
A new tomato NAC (NAM/ATAF1/2/CUC2) transcription factor, SlNAC4, functions as a positive regulator of fruit ripening and carotenoid accumulation.Plant Cell Physiol. 2014 Jan;55(1):119-35. doi: 10.1093/pcp/pct162. Epub 2013 Nov 20. Plant Cell Physiol. 2014. PMID: 24265273
-
The interplay between ABA/ethylene and NAC TFs in tomato fruit ripening: a review.Plant Mol Biol. 2021 Jun;106(3):223-238. doi: 10.1007/s11103-021-01128-w. Epub 2021 Feb 25. Plant Mol Biol. 2021. PMID: 33634368 Review.
-
Noncoding RNAs: functional regulatory factors in tomato fruit ripening.Theor Appl Genet. 2020 May;133(5):1753-1762. doi: 10.1007/s00122-020-03582-4. Epub 2020 Mar 24. Theor Appl Genet. 2020. PMID: 32211918 Review.
Cited by
-
MaNAC029 modulates ethylene biosynthesis and fruit quality and undergoes MaXB3-mediated proteasomal degradation during banana ripening.J Adv Res. 2023 Nov;53:33-47. doi: 10.1016/j.jare.2022.12.004. Epub 2022 Dec 15. J Adv Res. 2023. PMID: 36529351 Free PMC article.
-
A kiwifruit (Actinidia deliciosa) R2R3-MYB transcription factor modulates chlorophyll and carotenoid accumulation.New Phytol. 2019 Jan;221(1):309-325. doi: 10.1111/nph.15362. Epub 2018 Aug 1. New Phytol. 2019. PMID: 30067292 Free PMC article.
-
NAC61 regulates late- and post-ripening osmotic, oxidative, and biotic stress responses in grapevine.J Exp Bot. 2024 Apr 15;75(8):2330-2350. doi: 10.1093/jxb/erad507. J Exp Bot. 2024. PMID: 38159048 Free PMC article.
-
Molecular and Hormonal Mechanisms Regulating Fleshy Fruit Ripening.Cells. 2021 May 8;10(5):1136. doi: 10.3390/cells10051136. Cells. 2021. PMID: 34066675 Free PMC article. Review.
-
Transcription Factor CmNAC34 Regulated CmLCYB-Mediated β-Carotene Accumulation during Oriental Melon Fruit Ripening.Int J Mol Sci. 2022 Aug 29;23(17):9805. doi: 10.3390/ijms23179805. Int J Mol Sci. 2022. PMID: 36077205 Free PMC article.
References
-
- Costa F, Alba R, Schouten H, Soglio V, Gianfranceschi L, Serra S, Musacchi S, Sansavini S, Costa G, Fei ZJ, Giovannoni J. Use of homologous and heterologous gene expression profiling tools to characterize transcription dynamics during apple fruit maturation and ripening. BMC Plant Biol. 2010;10:229. doi: 10.1186/1471-2229-10-229. - DOI - PMC - PubMed
-
- Zouine M, Latché A, Rousseau C, Regad F, Pech JC, Philippot M, Bouzayen M, Delalande C, Frasse P, Schiex T, Noirot C, Bellec A, Klopp C, Berges H, Mariette J, Vautrin S, Causse M, Rothan C. The tomato genome sequence provides insights into fleshy fruit evolution. Nature. 2012;485:635–641. doi: 10.1038/nature11119. - DOI - PMC - PubMed
-
- Grierson D. Ethylene and the control of fruit ripening. In: Seymour GB, Giovannoni JJ, Tucker GA, Poole M, editors. Molecular Biology and Biochemistry of Fruit Ripening. Boston: Wiley In Press; 2013.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

