14-3-3 proteins participate in light signaling through association with PHYTOCHROME INTERACTING FACTORs
- PMID: 25501334
- PMCID: PMC4284738
- DOI: 10.3390/ijms151222801
14-3-3 proteins participate in light signaling through association with PHYTOCHROME INTERACTING FACTORs
Abstract
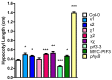
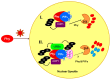
14-3-3 proteins are regulatory proteins found in all eukaryotes and are known to selectively interact with phosphorylated proteins to regulate physiological processes. Through an affinity purification screening, many light-related proteins were recovered as 14-3-3 candidate binding partners. Yeast two-hybrid analysis revealed that the 14-3-3 kappa isoform (14-3-3κ) could bind to PHYTOCHROME INTERACTING FACTOR3 (PIF3) and CONSTITUTIVE PHOTOMORPHOGENIC1 (COP1). Further analysis by in vitro pull-down assay confirmed the interaction between 14-3-3κ and PIF3. Interruption of putative phosphorylation sites on the 14-3-3 binding motifs of PIF3 was not sufficient to inhibit 14-3-3κ from binding or to disturb nuclear localization of PIF3. It was also indicated that 14-3-3κ could bind to other members of the PIF family, such as PIF1 and PIF6, but not to LONG HYPOCOTYL IN FAR-RED1 (HFR1). 14-3-3 mutants, as well as the PIF3 overexpressor, displayed longer hypocotyls, and a pif3 mutant displayed shorter hypocotyls than the wild-type in red light, suggesting that 14-3-3 proteins are positive regulators of photomorphogenesis and function antagonistically with PIF3. Consequently, our results indicate that 14-3-3 proteins bind to PIFs and initiate photomorphogenesis in response to a light signal.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

