Expression of major photosynthetic and salt-resistance genes in invasive reed lineages grown under elevated CO2 and temperature
- PMID: 25505541
- PMCID: PMC4242567
- DOI: 10.1002/ece3.1282
Expression of major photosynthetic and salt-resistance genes in invasive reed lineages grown under elevated CO2 and temperature
Abstract
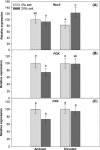
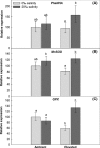
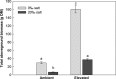
It is important to investigate the molecular causes of the variation in ecologically important traits to fully understand phenotypic responses to climate change. In the Mississippi River Delta, two distinct, sympatric invasive lineages of common reed (Phragmites australis) are known to differ in several ecophysiological characteristics and are expected to become more salt resistant due to increasing atmospheric CO2 and temperature. We investigated whether different patterns of gene expression can explain their ecophysiological differences and increased vigor under future climatic conditions. We compared the transcript abundance of photosynthetic genes of the Calvin cycle (Rubisco small subunit, RbcS; Phosphoglycerate kinase, PGK; Phosphoribulokinase, PRK), genes related with salt transport (Na(+)/H(+) antiporter, PhaNHA) and oxidative stress response genes (Manganese Superoxide dismutase, MnSOD; Glutathione peroxidase, GPX), and the total aboveground biomass production between two genotypes representing the two lineages. The two genotypes (Delta-type, Mediterranean lineage, and EU-type, Eurasian lineage) were grown under an ambient and a future climate scenario with simultaneously elevated CO2 and temperature, and under two different soil salinities (0‰ or 20‰). We found neither differences in the aboveground biomass production nor the transcript abundances of the two genotypes, but soil salinity significantly affected all the investigated parameters, often interacting with the climatic conditions. At 20‰ salinity, most genes were higher expressed in the future than in the ambient climatic conditions. Higher transcription of the genes suggests higher abundance of the protein they code for, and consequently increased photosynthate production, improved stress responses, and salt exclusion. Therefore, the higher expression of these genes most likely contributed to the significantly ameliorated salinity impact on the aboveground biomass production of both P. australis genotypes under elevated temperature and CO2. Although transcript abundances did not explain differences between the lineages, they correlated with the increased vigor of both lineages under anticipated future climatic conditions.
Keywords: Common reed; Delta-type; EU-type; Mississippi River Delta; Phragmites australis; US Gulf Coast.
Figures

Similar articles
-
Increased invasive potential of non-native Phragmites australis: elevated CO2 and temperature alleviate salinity effects on photosynthesis and growth.Glob Chang Biol. 2014 Feb;20(2):531-43. doi: 10.1111/gcb.12346. Glob Chang Biol. 2014. PMID: 23913622
-
Differentially expressed genes related to oxidoreductase activity and glutathione metabolism underlying the adaptation of Phragmites australis from the salt marsh in the Yellow River Delta, China.PeerJ. 2020 Oct 2;8:e10024. doi: 10.7717/peerj.10024. eCollection 2020. PeerJ. 2020. PMID: 33072439 Free PMC article.
-
Cosmopolitan Species As Models for Ecophysiological Responses to Global Change: The Common Reed Phragmites australis.Front Plant Sci. 2017 Nov 16;8:1833. doi: 10.3389/fpls.2017.01833. eCollection 2017. Front Plant Sci. 2017. PMID: 29250081 Free PMC article. Review.
-
Salt Water Exposure Exacerbates the Negative Response of Phragmites australis Haplotypes to Sea-Level Rise.Plants (Basel). 2024 Mar 21;13(6):906. doi: 10.3390/plants13060906. Plants (Basel). 2024. PMID: 38592938 Free PMC article.
-
Clonal integration promotes the growth of Phragmites australis populations in saline wetlands of the Yellow River Delta.Front Plant Sci. 2023 Jun 2;14:1162923. doi: 10.3389/fpls.2023.1162923. eCollection 2023. Front Plant Sci. 2023. PMID: 37332707 Free PMC article.
Cited by
-
How drought and ploidy level shape gene expression and DNA methylation in Phragmites australis.Plant Cell Rep. 2025 Aug 12;44(9):197. doi: 10.1007/s00299-025-03585-9. Plant Cell Rep. 2025. PMID: 40794332 Free PMC article.
-
Using Transcriptomics to Identify Differential Gene Expression in Response to Salinity among Australian Phragmites australis Clones.Front Plant Sci. 2016 Apr 13;7:432. doi: 10.3389/fpls.2016.00432. eCollection 2016. Front Plant Sci. 2016. PMID: 27148279 Free PMC article.
-
Comparative Proteomic Analysis of Two Manilkara Species Leaves Under NaCl Stress.Iran J Biotechnol. 2019 Sep 1;17(3):e2219. doi: 10.29252/ijb.2219. eCollection 2019 Sep. Iran J Biotechnol. 2019. PMID: 32195285 Free PMC article.
-
Discovery of Euryhaline Phycoerythrobilin-Containing Synechococcus and Its Mechanisms for Adaptation to Estuarine Environments.mSystems. 2020 Dec 15;5(6):e00842-20. doi: 10.1128/mSystems.00842-20. mSystems. 2020. PMID: 33323414 Free PMC article.
-
Choose Wisely: Great Variation among Genotypes of Promising Paludiculture Crop Phragmites australis.Plants (Basel). 2023 Feb 24;12(5):1045. doi: 10.3390/plants12051045. Plants (Basel). 2023. PMID: 36903906 Free PMC article.
References
-
- Achenbach L, Eller F, Nguyen LX, Brix H. Differences in salinity tolerance of genetically distinct Phragmites australis clones. AoB Plants. 2013;5:plt019. doi: 10.1093/aobpla/plt019. - DOI
-
- Ainsworth EA, Long SP. What have we learned from 15 years of free-air CO2 enrichment (FACE)? A meta-analytic review of the responses of photosynthesis, canopy. New Phytol. 2004;165:351–371. - PubMed
-
- Baerenfaller K, Grossmann J, Grobei MA, Hull R, Hirsch-Hoffmann M, Yalovsky S, et al. Genome-scale proteomics reveals Arabidopsis thaliana gene models and proteome dynamics. Science. 2008;320:938–941. - PubMed
-
- Barnola JM, Anklin M, Porcheron J, Raynaud D, Schwander J, Stauffer B. CO2 evolution during the last millennium as recorded by Antarctic and Greenland ice. Tellus Ser. B. 1995;47B:264–272.
LinkOut - more resources
Full Text Sources
Other Literature Sources

