Composition, roles, and regulation of cullin-based ubiquitin e3 ligases
- PMID: 25505853
- PMCID: PMC4262284
- DOI: 10.1199/tab.0175
Composition, roles, and regulation of cullin-based ubiquitin e3 ligases
Abstract
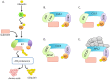
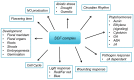
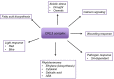
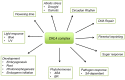
Due to their sessile nature, plants depend on flexible regulatory systems that allow them to adequately regulate developmental and physiological processes in context with environmental cues. The ubiquitin proteasome pathway, which targets a great number of proteins for degradation, is cellular tool that provides the necessary flexibility to accomplish this task. Ubiquitin E3 ligases provide the needed specificity to the pathway by selectively binding to particular substrates and facilitating their ubiquitylation. The largest group of E3 ligases known in plants is represented by CULLIN-REALLY INTERESTING NEW GENE (RING) E3 ligases (CRLs). In recent years, a great amount of knowledge has been generated to reveal the critical roles of these enzymes across all aspects of plant life. This review provides an overview of the different classes of CRLs in plants, their specific complex compositions, the variety of biological processes they control, and the regulatory steps that can affect their activities.
Figures
Similar articles
-
Cullin 5-RING E3 ubiquitin ligases, new therapeutic targets?Biochimie. 2016 Mar;122:339-47. doi: 10.1016/j.biochi.2015.08.003. Epub 2015 Aug 4. Biochimie. 2016. PMID: 26253693 Review.
-
Regulation of cullin RING ligases.Annu Rev Plant Biol. 2008;59:467-89. doi: 10.1146/annurev.arplant.58.032806.104011. Annu Rev Plant Biol. 2008. PMID: 18444905 Review.
-
Introduction.Adv Exp Med Biol. 2020;1217:1-8. doi: 10.1007/978-981-15-1025-0_1. Adv Exp Med Biol. 2020. PMID: 31898218
-
Cullin 3, a cellular scripter of the non-proteolytic ubiquitin code.Semin Cell Dev Biol. 2019 Sep;93:100-110. doi: 10.1016/j.semcdb.2018.12.007. Epub 2018 Dec 28. Semin Cell Dev Biol. 2019. PMID: 30586619 Review.
-
The cullin-RING ubiquitin-protein ligases.Annu Rev Plant Biol. 2011;62:299-334. doi: 10.1146/annurev-arplant-042809-112256. Annu Rev Plant Biol. 2011. PMID: 21370976 Review.
Cited by
-
Mechanism of cytoplasmic mRNA translation.Arabidopsis Book. 2015 Apr 24;13:e0176. doi: 10.1199/tab.0176. eCollection 2015. Arabidopsis Book. 2015. PMID: 26019692 Free PMC article.
-
Aluminum resistance in plants: A critical review focusing on STOP1.Plant Commun. 2025 Feb 10;6(2):101200. doi: 10.1016/j.xplc.2024.101200. Epub 2024 Dec 2. Plant Commun. 2025. PMID: 39628052 Free PMC article. Review.
-
Immunoprecipitation of Cullin-Ring Ligases (CRLs) in Arabidopsis thaliana Seedlings.Methods Mol Biol. 2023;2581:31-42. doi: 10.1007/978-1-0716-2784-6_3. Methods Mol Biol. 2023. PMID: 36413308
-
The ubiquitination machinery of the ubiquitin system.Arabidopsis Book. 2014 Oct 6;12:e0174. doi: 10.1199/tab.0174. eCollection 2014. Arabidopsis Book. 2014. PMID: 25320573 Free PMC article.
-
No Time to Waste: Transcriptome Study Reveals that Drought Tolerance in Barley May Be Attributed to Stressed-Like Expression Patterns that Exist before the Occurrence of Stress.Front Plant Sci. 2018 Jan 9;8:2212. doi: 10.3389/fpls.2017.02212. eCollection 2017. Front Plant Sci. 2018. PMID: 29375595 Free PMC article.
References
-
- Al Khateeb W.M., Schroeder D.F. Overexpression of Arabidopsis damaged DNA binding protein 1A (DDB1A) enhances UV tolerance. Plant Mol. Biol. 2009;70:371–383. - PubMed
-
- Alonso-Peral M.M., Candela H., del Pozo J.C., Martinez-Laborda A., Ponce M.R., Micol J.L. The HVE/CAND1 gene is required for the early patterning of leaf venation in Arabidopsis. Development. 2006;133:3755–3766. - PubMed
-
- Alonso J.M., Hirayama T., Roman G., Nourizadeh S., Ecker J.R. EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science. 1999;284:2148–2152. - PubMed
-
- Anand A., Rojas C.M., Tang Y., Mysore K.S. Several components of SKP1/Cullin/F-box E3 ubiquitin ligase complex and associated factors play a role in Agrobacterium-mediated plant transformation. New Phytol. 2012;195:203–216. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
