Mutations within A 35 amino acid region of P6 influence self-association, inclusion body formation, and Caulimovirus infectivity
- PMID: 25506670
- PMCID: PMC4323857
- DOI: 10.1016/j.virol.2014.11.017
Mutations within A 35 amino acid region of P6 influence self-association, inclusion body formation, and Caulimovirus infectivity
Abstract
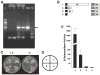
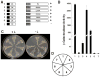
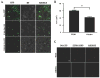
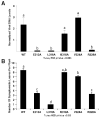
Cauliflower mosaic virus gene VI product (P6) is an essential protein that forms cytoplasmic, inclusion bodies (IBs). P6 contains four regions involved in self-association, termed D1-D4. D3 binds to D1, along with D4 and contains a spacer region (termed D3b) between two RNA-binding domains. Here we show D3b binds full-length P6 along with D1 and D4. Full-length P6s harboring single amino acid substitutions within D3b showed reduced binding to both D1 and D4. Full-length P6s containing D3b mutations and fused with green fluorescent protein formed inclusion-like bodies (IL-Bs) when expressed in Nicotiana benthamiana leaves. However, mutant P6s with reduced binding to D1 and D4, showed smaller IL-Bs, than wild type. Likewise, viruses containing these mutations showed a decrease in inoculated leaf viral DNA levels and reduced efficiency of systemic infection. These data suggest that mutations influencing P6 self-association alter IB formation and reduce virus infection.
Keywords: CaMV; Gene VI product; Inclusion bodies; TAV.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Formation of large viroplasms and virulence of Cauliflower mosaic virus in turnip plants depend on the N-terminal EKI sequence of viral protein TAV.PLoS One. 2017 Dec 18;12(12):e0189062. doi: 10.1371/journal.pone.0189062. eCollection 2017. PLoS One. 2017. PMID: 29253877 Free PMC article.
-
Setting Up Shop: The Formation and Function of the Viral Factories of Cauliflower mosaic virus.Front Plant Sci. 2017 Oct 30;8:1832. doi: 10.3389/fpls.2017.01832. eCollection 2017. Front Plant Sci. 2017. PMID: 29163571 Free PMC article. Review.
-
The open reading frame VI product of Cauliflower mosaic virus is a nucleocytoplasmic protein: its N terminus mediates its nuclear export and formation of electron-dense viroplasms.Plant Cell. 2005 Mar;17(3):927-43. doi: 10.1105/tpc.104.029017. Plant Cell. 2005. PMID: 15746075 Free PMC article.
-
Multiple domains within the Cauliflower mosaic virus gene VI product interact with the full-length protein.Mol Plant Microbe Interact. 2002 Oct;15(10):1050-7. doi: 10.1094/MPMI.2002.15.10.1050. Mol Plant Microbe Interact. 2002. PMID: 12437303
-
A model for intracellular movement of Cauliflower mosaic virus: the concept of the mobile virion factory.J Exp Bot. 2016 Mar;67(7):2039-48. doi: 10.1093/jxb/erv520. Epub 2015 Dec 18. J Exp Bot. 2016. PMID: 26687180 Review.
Cited by
-
Formation of large viroplasms and virulence of Cauliflower mosaic virus in turnip plants depend on the N-terminal EKI sequence of viral protein TAV.PLoS One. 2017 Dec 18;12(12):e0189062. doi: 10.1371/journal.pone.0189062. eCollection 2017. PLoS One. 2017. PMID: 29253877 Free PMC article.
-
Setting Up Shop: The Formation and Function of the Viral Factories of Cauliflower mosaic virus.Front Plant Sci. 2017 Oct 30;8:1832. doi: 10.3389/fpls.2017.01832. eCollection 2017. Front Plant Sci. 2017. PMID: 29163571 Free PMC article. Review.
References
-
- Acosta-Leal R, Duffy S, Xiong Z, Hammond RW, Elena SF. Advances in plant virus evolution: translating evolutionary insights into better disease management. Phytopathology. 2011;101:1136–1148. - PubMed
-
- Agama K, Beach S, Schoelz J, Leisner SM. the 5′ third of Cauliflower mosaic virus gene VI conditions resistance breakage in Arabidopsis ecotype Tsu-0. Phytopathology. 2002;92:190–196. - PubMed
-
- Anderson EJ, Trese AT, Sehgal OP, Schoelz JE. Characterization of a chimeric Cauliflower mosaic virus isolate that is more severe and accumulates to higher concentrations the either of the strains from which it was derived. Mol Plant-Microbe Interact. 1992;5:48–54.
-
- Angel CA, Lutz L, Yang X, Rodriguez A, Adair A, Zhang Y, Leisner SM, Nelson RS, Schoelz JE. The P6 protein of Cauliflower mosaic virus interacts with CHUP1, a plant protein which moves chloroplasts on actin microfilaments. Virology. 2013;443:363–374. - PubMed
-
- Armour SL, Melcher U, Pirone TP, Lyttle DJ, Essenberg RC. Helper component for aphid transmission encoded by region II of Cauliflower mosaic virus DNA. Virology. 1983;129:25–30. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

