In silico selection of therapeutic antibodies for development: viscosity, clearance, and chemical stability
- PMID: 25512516
- PMCID: PMC4284567
- DOI: 10.1073/pnas.1421779112
In silico selection of therapeutic antibodies for development: viscosity, clearance, and chemical stability
Erratum in
-
Correction for Sharma et al., In silico selection of therapeutic antibodies for development: Viscosity, clearance, and chemical stability.Proc Natl Acad Sci U S A. 2015 Dec 1;112(48):E6719. doi: 10.1073/pnas.1519997112. Epub 2015 Nov 23. Proc Natl Acad Sci U S A. 2015. PMID: 26598676 Free PMC article. No abstract available.
Abstract
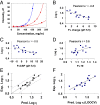
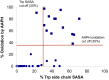
For mAbs to be viable therapeutics, they must be formulated to have low viscosity, be chemically stable, and have normal in vivo clearance rates. We explored these properties by observing correlations of up to 60 different antibodies of the IgG1 isotype. Unexpectedly, we observe significant correlations with simple physical properties obtainable from antibody sequences and by molecular dynamics simulations of individual antibody molecules. mAbs viscosities increase strongly with hydrophobicity and charge dipole distribution and decrease with net charge. Fast clearance correlates with high hydrophobicities of certain complementarity determining regions and with high positive or high negative net charge. Chemical degradation from tryptophan oxidation correlates with the average solvent exposure time of tryptophan residues. Aspartic acid isomerization rates can be predicted from solvent exposure and flexibility as determined by molecular dynamics simulations. These studies should aid in more rapid screening and selection of mAb candidates during early discovery.
Keywords: degradation; monoclonal antibodies; pharmacokinetics; prediction; viscosity.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Eisenstein M. Something new under the skin. Nat Biotechnol. 2011;29(2):107–109. - PubMed
-
- Shire SJ, Shahrokh Z, Liu J. Challenges in the development of high protein concentration formulations. J Pharm Sci. 2004;93(6):1390–1402. - PubMed
-
- Wang W, Wang EQ, Balthasar JP. Monoclonal antibody pharmacokinetics and pharmacodynamics. Clin Pharmacol Ther. 2008;84(5):548–558. - PubMed
-
- Buck PM, et al. Computational methods to predict therapeutic protein aggregation. Therapeutic Proteins: Methods and Protocols. 2012 eds Voynov V, Caravella JA (Springer, New York), Vol 899, pp 425–451. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

