Katanin p80 regulates human cortical development by limiting centriole and cilia number
- PMID: 25521379
- PMCID: PMC4485387
- DOI: 10.1016/j.neuron.2014.12.017
Katanin p80 regulates human cortical development by limiting centriole and cilia number
Abstract
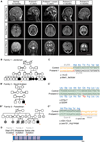
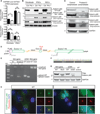
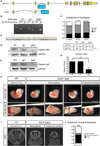
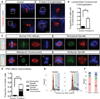
Katanin is a microtubule-severing complex whose catalytic activities are well characterized, but whose in vivo functions are incompletely understood. Human mutations in KATNB1, which encodes the noncatalytic regulatory p80 subunit of katanin, cause severe microlissencephaly. Loss of Katnb1 in mice confirms essential roles in neurogenesis and cell survival, while loss of zebrafish katnb1 reveals specific roles for katnin p80 in early and late developmental stages. Surprisingly, Katnb1 null mutant mouse embryos display hallmarks of aberrant Sonic hedgehog signaling, including holoprosencephaly. KATNB1-deficient human cells show defective proliferation and spindle structure, while Katnb1 null fibroblasts also demonstrate a remarkable excess of centrioles, with supernumerary cilia but deficient Hedgehog signaling. Our results reveal unexpected functions for KATNB1 in regulating overall centriole, mother centriole, and cilia number, and as an essential gene for normal Hedgehog signaling during neocortical development.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures



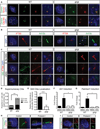
References
-
- Arquint C, Nigg EA. STIL microcephaly mutations interfere with APC/C-mediated degradation and cause centriole amplification. Curr. Biol. 2014;24:351–360. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM095941/GM/NIGMS NIH HHS/United States
- RC2MH089952/MH/NIMH NIH HHS/United States
- T32HL007731/HL/NHLBI NIH HHS/United States
- RC2 MH089952/MH/NIMH NIH HHS/United States
- T32 GM007753/GM/NIGMS NIH HHS/United States
- R01NS035129/NS/NINDS NIH HHS/United States
- P30 HD018655/HD/NICHD NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- R01 NS035129/NS/NINDS NIH HHS/United States
- T32 HL007731/HL/NHLBI NIH HHS/United States
- R01 AR054396/AR/NIAMS NIH HHS/United States
- 1U01DE024434-01/DE/NIDCR NIH HHS/United States
- U01 DE024434/DE/NIDCR NIH HHS/United States
- R01AR054396/AR/NIAMS NIH HHS/United States
- T32GM007753/GM/NIGMS NIH HHS/United States
- R01GM095941/GM/NIGMS NIH HHS/United States
- F31NS083111/NS/NINDS NIH HHS/United States
- F31 NS083111/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

