Comparative transcriptome analyses reveal core parasitism genes and suggest gene duplication and repurposing as sources of structural novelty
- PMID: 25534030
- PMCID: PMC4327159
- DOI: 10.1093/molbev/msu343
Comparative transcriptome analyses reveal core parasitism genes and suggest gene duplication and repurposing as sources of structural novelty
Abstract
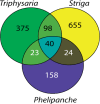
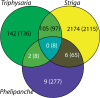
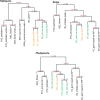
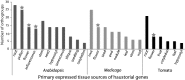
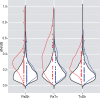
The origin of novel traits is recognized as an important process underlying many major evolutionary radiations. We studied the genetic basis for the evolution of haustoria, the novel feeding organs of parasitic flowering plants, using comparative transcriptome sequencing in three species of Orobanchaceae. Around 180 genes are upregulated during haustorial development following host attachment in at least two species, and these are enriched in proteases, cell wall modifying enzymes, and extracellular secretion proteins. Additionally, about 100 shared genes are upregulated in response to haustorium inducing factors prior to host attachment. Collectively, we refer to these newly identified genes as putative "parasitism genes." Most of these parasitism genes are derived from gene duplications in a common ancestor of Orobanchaceae and Mimulus guttatus, a related nonparasitic plant. Additionally, the signature of relaxed purifying selection and/or adaptive evolution at specific sites was detected in many haustorial genes, and may play an important role in parasite evolution. Comparative analysis of gene expression patterns in parasitic and nonparasitic angiosperms suggests that parasitism genes are derived primarily from root and floral tissues, but with some genes co-opted from other tissues. Gene duplication, often taking place in a nonparasitic ancestor of Orobanchaceae, followed by regulatory neofunctionalization, was an important process in the origin of parasitic haustoria.
Keywords: Orobanchaceae; gene duplication; novel traits; parasitism; protease; transcriptome.
© The Author 2014. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures


References
-
- Aber M, Fer A, Salle G. Transfer of organic substances from the host plant Vicia faba to the parasite Orobanche renata Forsk. Z Pflanzenphysiol. 1983;112:297–308.
-
- Aly R, Cholakh H, Joel DM, Leibman D, Steinitz B, Zelcer A, Naglis A, Yarden O, Gal-On A. Gene silencing of mannose 6-phosphate reductase in the parasitic weed Orobanche aegyptiaca through the production of homologous dsRNA sequences in the host plant. Plant Biotechnol J. 2009;7:487–498. - PubMed
-
- Aly R, Hamamouch N, Abu-Nassar J, et al. Movement of protein and macromolecules between host plants and the parasitic weed Phelipanche aegyptiaca Pers. Plant Cell Rep. 2011;30:2233–2241. - PubMed
-
- Amborella Genome Project. The Amborella genome and the evolution of flowering plants. Science. 2013;342:1241089. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

