DNA ligase III acts as a DNA strand break sensor in the cellular orchestration of DNA strand break repair
- PMID: 25539916
- PMCID: PMC4333375
- DOI: 10.1093/nar/gku1307
DNA ligase III acts as a DNA strand break sensor in the cellular orchestration of DNA strand break repair
Abstract
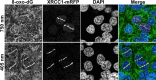
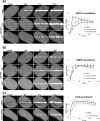
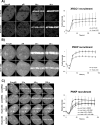
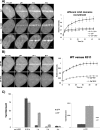
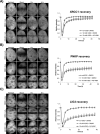
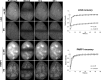
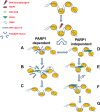
In the current model of DNA SSBR, PARP1 is regarded as the sensor of single-strand breaks (SSBs). However, biochemical studies have implicated LIG3 as another possible SSB sensor. Using a laser micro-irradiation protocol that predominantly generates SSBs, we were able to demonstrate that PARP1 is dispensable for the accumulation of different single-strand break repair (SSBR) proteins at sites of DNA damage in live cells. Furthermore, we show in live cells for the first time that LIG3 plays a role in mediating the accumulation of the SSBR proteins XRCC1 and PNKP at sites of DNA damage. Importantly, the accumulation of LIG3 at sites of DNA damage did not require the BRCT domain-mediated interaction with XRCC1. We were able to show that the N-terminal ZnF domain of LIG3 plays a key role in the enzyme's SSB sensing function. Finally, we provide cellular evidence that LIG3 and not PARP1 acts as the sensor for DNA damage caused by the topoisomerase I inhibitor, irinotecan. Our results support the existence of a second damage-sensing mechanism in SSBR involving the detection of nicks in the genome by LIG3.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




Similar articles
-
Overlapping roles for PARP1 and PARP2 in the recruitment of endogenous XRCC1 and PNKP into oxidized chromatin.Nucleic Acids Res. 2017 Mar 17;45(5):2546-2557. doi: 10.1093/nar/gkw1246. Nucleic Acids Res. 2017. PMID: 27965414 Free PMC article.
-
Disconnecting XRCC1 and DNA ligase III.Cell Cycle. 2011 Jul 15;10(14):2269-75. doi: 10.4161/cc.10.14.16495. Epub 2011 Jul 15. Cell Cycle. 2011. PMID: 21636980 Free PMC article.
-
Physical and functional interaction between DNA ligase IIIalpha and poly(ADP-Ribose) polymerase 1 in DNA single-strand break repair.Mol Cell Biol. 2003 Aug;23(16):5919-27. doi: 10.1128/MCB.23.16.5919-5927.2003. Mol Cell Biol. 2003. PMID: 12897160 Free PMC article.
-
Structure and function of the DNA ligases encoded by the mammalian LIG3 gene.Gene. 2013 Dec 1;531(2):150-7. doi: 10.1016/j.gene.2013.08.061. Epub 2013 Sep 5. Gene. 2013. PMID: 24013086 Free PMC article. Review.
-
Joining of DNA breaks- interplay between DNA ligases and poly (ADP-ribose) polymerases.DNA Repair (Amst). 2025 May;149:103843. doi: 10.1016/j.dnarep.2025.103843. Epub 2025 May 2. DNA Repair (Amst). 2025. PMID: 40347914 Review.
Cited by
-
Sensitization of colorectal cancer to irinotecan therapy by PARP inhibitor rucaparib.Invest New Drugs. 2019 Oct;37(5):948-960. doi: 10.1007/s10637-018-00717-9. Epub 2019 Jan 5. Invest New Drugs. 2019. PMID: 30612311
-
XRCC1-mediated repair of strand breaks independent of PNKP binding.DNA Repair (Amst). 2017 Dec;60:52-63. doi: 10.1016/j.dnarep.2017.10.007. Epub 2017 Oct 19. DNA Repair (Amst). 2017. PMID: 29100039 Free PMC article.
-
The small GTPase RhoU lays downstream of JAK/STAT signaling and mediates cell migration in multiple myeloma.Blood Cancer J. 2018 Feb 13;8(2):20. doi: 10.1038/s41408-018-0053-z. Blood Cancer J. 2018. PMID: 29440639 Free PMC article.
-
PARP1 condensates differentially partition DNA repair proteins and enhance DNA ligation.EMBO Rep. 2024 Dec;25(12):5635-5666. doi: 10.1038/s44319-024-00285-5. Epub 2024 Nov 4. EMBO Rep. 2024. PMID: 39496836 Free PMC article.
-
The zinc finger of DNA Ligase 3α binds to nucleosomes via an arginine anchor.Res Sq [Preprint]. 2025 Mar 10:rs.3.rs-6033068. doi: 10.21203/rs.3.rs-6033068/v1. Res Sq. 2025. PMID: 40162218 Free PMC article. Preprint.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

