Global mapping of herpesvirus-host protein complexes reveals a transcription strategy for late genes
- PMID: 25544563
- PMCID: PMC4305015
- DOI: 10.1016/j.molcel.2014.11.026
Global mapping of herpesvirus-host protein complexes reveals a transcription strategy for late genes
Abstract
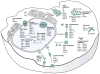
Mapping host-pathogen interactions has proven instrumental for understanding how viruses manipulate host machinery and how numerous cellular processes are regulated. DNA viruses such as herpesviruses have relatively large coding capacity and thus can target an extensive network of cellular proteins. To identify the host proteins hijacked by this pathogen, we systematically affinity tagged and purified all 89 proteins of Kaposi's sarcoma-associated herpesvirus (KSHV) from human cells. Mass spectrometry of this material identified over 500 virus-host interactions. KSHV causes AIDS-associated cancers, and its interaction network is enriched for proteins linked to cancer and overlaps with proteins that are also targeted by HIV-1. We found that the conserved KSHV protein ORF24 binds to RNA polymerase II and brings it to viral late promoters by mimicking and replacing cellular TATA-box-binding protein (TBP). This is required for herpesviral late gene expression, a complex and poorly understood phase of the viral lifecycle.
Copyright © 2015 Elsevier Inc. All rights reserved.
Figures






References
-
- Bushnell DA, Westover KD, Davis RE, Kornberg RD. Structural basis of transcription: an RNA polymerase II-TFIIB cocrystal at 4.5 Angstroms. Science. 2004;303:983–988. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01HG007644/HG/NHGRI NIH HHS/United States
- U54 AI081680/AI/NIAID NIH HHS/United States
- R01 CA160556/CA/NCI NIH HHS/United States
- F31 CA180609-01/CA/NCI NIH HHS/United States
- F31 CA180609/CA/NCI NIH HHS/United States
- CA160556/CA/NCI NIH HHS/United States
- P50 GM081879/GM/NIGMS NIH HHS/United States
- R01 HG007644/HG/NHGRI NIH HHS/United States
- P50 GM082250/GM/NIGMS NIH HHS/United States
- R01 CA136367/CA/NCI NIH HHS/United States
- P01 CA177322/CA/NCI NIH HHS/United States
- P01 AI091575/AI/NIAID NIH HHS/United States
- P01 AI090935/AI/NIAID NIH HHS/United States
- CA136367/CA/NCI NIH HHS/United States
- P30 AI027763/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

