Surveying the global virome: identification and characterization of HCV-related animal hepaciviruses
- PMID: 25545071
- PMCID: PMC5081135
- DOI: 10.1016/j.antiviral.2014.12.014
Surveying the global virome: identification and characterization of HCV-related animal hepaciviruses
Abstract
Recent advances in sequencing technologies have greatly enhanced our abilities to identify novel microbial sequences. Thus, our understanding of the global virome and the virome of specific host species in particular is rapidly expanding. Identification of animal viruses is important for understanding animal disease, the origin and evolution of human viruses, as well as zoonotic reservoirs for emerging infections. Although the human hepacivirus, hepatitis C virus (HCV), was identified 25years ago, its origin has remained elusive. In 2011, the first HCV homolog was reported in dogs but subsequent studies showed the virus to be widely distributed in horses. This indicated a wider hepacivirus host range and paved the way for identification of rodent, bat and non-human primate hepaciviruses. The equine non-primate hepacivirus (NPHV) remains the closest relative of HCV and is so far the best characterized. Identification and characterization of novel hepaciviruses may in addition lead to development of tractable animal models to study HCV persistence, immune responses and pathogenesis. This could be particular important, given the current shortage of immunocompetent models for robust HCV infection. Much remains to be learned on the novel hepaciviruses, including their association with disease, and thereby how relevant they will become as HCV model systems and for studies of animal disease. This review discusses how virome analysis led to identification of novel hepaci- and pegiviruses, their genetic relationship and characterization and the potential use of animal hepaciviruses as models to study hepaciviral infection, immunity and pathogenesis. This article forms part of a symposium in Antiviral Research on "Hepatitis C: Next steps toward global eradication."
Copyright © 2015 Elsevier B.V. All rights reserved.
Figures
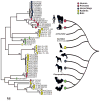
References
-
- Anthony SJ, Epstein JH, Murray KA, Navarrete-Macias I, Zambrana-Torrelio CM, Solovyov A, Ojeda-Flores R, Arrigo NC, Islam A, Ali Khan S, Hosseini P, Bogich TL, Olival KJ, Sanchez-Leon MD, Karesh WB, Goldstein T, Luby SP, Morse SS, Mazet JA, Daszak P, Lipkin WI. A strategy to estimate unknown viral diversity in mammals. MBio. 2013;4:e00598–00513. - PMC - PubMed
-
- Bukh J. Animal models for the study of hepatitis C virus infection and related liver disease. Gastroenterology. 2012;142:1279–1287. e1273. - PubMed
-
- Bukh J, Apgar CL, Yanagi M. Toward a surrogate model for hepatitis C virus: An infectious molecular clone of the GB virus-B hepatitis agent. Virology. 1999;262:470–478. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

