New insight into secreted ribonuclease structure: binase is a natural dimer
- PMID: 25551440
- PMCID: PMC4281067
- DOI: 10.1371/journal.pone.0115818
New insight into secreted ribonuclease structure: binase is a natural dimer
Abstract
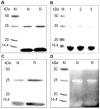
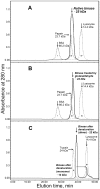
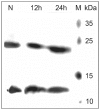
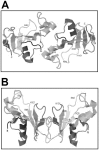
The biological effects of ribonucleases (RNases), such as the control of the blood vessels growth, the toxicity towards tumour cells and antiviral activity, require a detailed explanation. One of the most intriguing properties of RNases which can contribute to their biological effects is the ability to form dimers, which facilitates efficient RNA hydrolysis and the evasion of ribonuclease inhibitor. Dimeric forms of microbial RNase binase secreted by Bacillus pumilus (former B. intermedius) have only been found in crystals to date. Our study is the first report directly confirming the existence of binase dimers in solution and under natural conditions of enzyme biosynthesis and secretion by bacilli. Using different variants of gel electrophoresis, immunoblotting, size-exclusion chromatography and mass-spectrometry, we revealed that binase is a stable natural dimer with high catalytic activity.
Conflict of interest statement
Figures

References
-
- Shlyakhovenko VA (2009) Ribonucleases in tumor growth. Exp Oncol 31:127–133. - PubMed
-
- Cook GM, Robson JR, Frampton RA, McKenzie J, Przybilski R, et al. (2013) Ribonucleases in bacterial toxin-antitoxin systems. Biochim Biophys Acta 1829:523–531. - PubMed
-
- Bertram R, Schuster CF (2014) Post-transcriptional regulation of gene expression in bacterial pathogens by toxin-antitoxin systems. Front Cell Infect Microbiol 4 doi:10.3389/fcimb.2014.00006. - DOI - PMC - PubMed
-
- Ulyanova V, Vershinina V, Ilinskaya O (2011) Barnase and binase: twins with distinct fates. FEBS J 278:3633–3643. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

