Proteome scale comparative modeling for conserved drug and vaccine targets identification in Corynebacterium pseudotuberculosis
- PMID: 25573232
- PMCID: PMC4243142
- DOI: 10.1186/1471-2164-15-S7-S3
Proteome scale comparative modeling for conserved drug and vaccine targets identification in Corynebacterium pseudotuberculosis
Abstract
Corynebacterium pseudotuberculosis (Cp) is a pathogenic bacterium that causes caseous lymphadenitis (CLA), ulcerative lymphangitis, mastitis, and edematous to a broad spectrum of hosts, including ruminants, thereby threatening economic and dairy industries worldwide. Currently there is no effective drug or vaccine available against Cp. To identify new targets, we adopted a novel integrative strategy, which began with the prediction of the modelome (tridimensional protein structures for the proteome of an organism, generated through comparative modeling) for 15 previously sequenced C. pseudotuberculosis strains. This pan-modelomics approach identified a set of 331 conserved proteins having 95-100% intra-species sequence similarity. Next, we combined subtractive proteomics and modelomics to reveal a set of 10 Cp proteins, which may be essential for the bacteria. Of these, 4 proteins (tcsR, mtrA, nrdI, and ispH) were essential and non-host homologs (considering man, horse, cow and sheep as hosts) and satisfied all criteria of being putative targets. Additionally, we subjected these 4 proteins to virtual screening of a drug-like compound library. In all cases, molecules predicted to form favorable interactions and which showed high complementarity to the target were found among the top ranking compounds. The remaining 6 essential proteins (adk, gapA, glyA, fumC, gnd, and aspA) have homologs in the host proteomes. Their active site cavities were compared to the respective cavities in host proteins. We propose that some of these proteins can be selectively targeted using structure-based drug design approaches (SBDD). Our results facilitate the selection of C. pseudotuberculosis putative proteins for developing broad-spectrum novel drugs and vaccines. A few of the targets identified here have been validated in other microorganisms, suggesting that our modelome strategy is effective and can also be applicable to other pathogens.
Figures
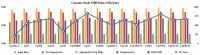
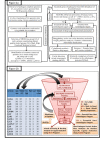
References
-
- Dorella FA, Pacheco LG, Oliveira SC, Miyoshi A, Azevedo V. Corynebacterium pseudotuberculosis: microbiology, biochemical properties, pathogenesis and molecular studies of virulence. Veterinary research. 2006;37(2):201–218. - PubMed
-
- Soares SC, Trost E, Ramos RT, Carneiro AR, Santos AR, Pinto AC, Barbosa E, Aburjaile F, Ali A, Diniz CA, Genome sequence of Corynebacterium pseudotuberculosis biovar equi strain 258 and prediction of antigenic targets to improve biotechnological vaccine production. Journal of biotechnology. 2012. - PubMed
-
- Williamson LH. Caseous lymphadenitis in small ruminants. Vet Clin North Am Food Anim Pract. 2001;17(2):359–371. vii. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

