Unity in diversity: shared mechanism of entry among paramyxoviruses
- PMID: 25595799
- PMCID: PMC4369139
- DOI: 10.1016/bs.pmbts.2014.10.001
Unity in diversity: shared mechanism of entry among paramyxoviruses
Abstract
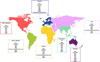
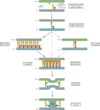
The Paramyxoviridae family includes many viruses that are pathogenic in humans, including parainfluenza viruses, measles virus, respiratory syncytial virus, and the emerging zoonotic Henipaviruses. No effective treatments are currently available for these viruses, and there is a need for efficient antiviral therapies. Paramyxoviruses enter the target cell by binding to a cell surface receptor and then fusing the viral envelope with the target cell membrane, allowing the release of the viral genome into the cytoplasm. Blockage of these crucial steps prevents infection and disease. Binding and fusion are driven by two virus-encoded glycoproteins, the receptor-binding protein and the fusion protein, that together form the viral "fusion machinery." The development of efficient antiviral drugs requires a deeper understanding of the mechanism of action of the Paramyxoviridae fusion machinery, which is still controversial. Here, we review recent structural and functional data on these proteins and the current understanding of the mechanism of the paramyxovirus cell entry process.
Keywords: Fusion machinery; Fusion protein; Paramyxoviridae; Receptor-binding protein; Viral entry.
© 2015 Elsevier Inc. All rights reserved.
Figures






References
-
- Enders G. Paramyxoviruses. In: Baron S, editor. Medical Microbiology. 4th ed. Galveston (TX): University of Texas Medical Branch at Galveston; 1996. [Accessed April 11, 2014]. Available at: http://www.ncbi.nlm.nih.gov/books/NBK8461/
-
- Falk K, Batts WN, Kvellestad A, Kurath G, Wiik-Nielsen J, Winton JR. Molecular characterisation of Atlantic salmon paramyxovirus (ASPV): a novel paramyxovirus associated with proliferative gill inflammation. Virus Res. 2008;133(2):218–227. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

