Epigenomic analysis of the HOX gene loci reveals mechanisms that may control canonical expression patterns in AML and normal hematopoietic cells
- PMID: 25600023
- PMCID: PMC4456213
- DOI: 10.1038/leu.2015.6
Epigenomic analysis of the HOX gene loci reveals mechanisms that may control canonical expression patterns in AML and normal hematopoietic cells
Abstract
HOX genes are highly expressed in many acute myeloid leukemia (AML) samples, but the patterns of expression and associated regulatory mechanisms are not clearly understood. We analyzed RNA sequencing data from 179 primary AML samples and normal hematopoietic cells to understand the range of expression patterns in normal versus leukemic cells. HOX expression in AML was restricted to specific genes in the HOXA or HOXB loci, and was highly correlated with recurrent cytogenetic abnormalities. However, the majority of samples expressed a canonical set of HOXA and HOXB genes that was nearly identical to the expression signature of normal hematopoietic stem/progenitor cells. Transcriptional profiles at the HOX loci were similar between normal cells and AML samples, and involved bidirectional transcription at the center of each gene cluster. Epigenetic analysis of a subset of AML samples also identified common regions of chromatin accessibility in AML samples and normal CD34(+) cells that displayed differences in methylation depending on HOX expression patterns. These data provide an integrated epigenetic view of the HOX gene loci in primary AML samples, and suggest that HOX expression in most AML samples represents a normal stem cell program that is controlled by epigenetic mechanisms at specific regulatory elements.
Conflict of interest statement
The authors have no conflicts of interest relating to the work described in this manuscript.
Figures
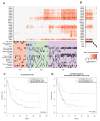




References
-
- Pineault N, Helgason CD, Lawrence HJ, Humphries RK. Differential expression of Hox, Meis1, and Pbx1 genes in primitive cells throughout murine hematopoietic ontogeny. Experimental Hematology. 2002 Jan;30(1):49–57. - PubMed
-
- Giampaolo A, Sterpetti P, Bulgarini D, Samoggia P, Pelosi E, Valtieri M, et al. Key functional role and lineage-specific expression of selected HOXB genes in purified hematopoietic progenitor differentiation. Blood. 1994 Dec 1;84(11):3637–47. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

